Integrated Bioinformatics and Experimental Analysis of KIFs in Ovarian Cancer Reveals Mitotic Drivers and Germline Variant Associations
Download
Abstract
Background: Ovarian cancer is the deadliest gynecologic malignancy, yet its genomic drivers remain incompletely understood.
Methods: We integrated multi-omics data from TCGA-OV (n = 379), GTEx normals (n = 88), and two GEO cohorts (GSE26712, GSE18520) to analyze differential expression, CNV, pathway enrichment, and protein–protein interactions. Germline variants were assessed through eQTL mapping and functional annotation. KIF17 expression and rs13375609 genotypes were experimentally validated using qRT-PCR and T-ARMS PCR in an independent cohort (12 tumors, 10 normals).
Results: We identified ~3,200 dysregulated genes, with mitotic kinesins (KIF11, KIF17, KIF18A, KIF20A) markedly upregulated and frequently amplified. High KIF11 or KIF14 expression correlated with reduced overall survival. GSEA indicated strong enrichment of mitotic spindle and G2/M checkpoint pathways, while PPI analysis identified KIF11 and KIF17 as central mitotic hubs. Two KIF17 variants (rs2297299, rs13375609) showed significant eQTL effects. Experimental validation confirmed elevated KIF17 expression and higher frequency of the rs13375609 A allele in ovarian cancer (27% vs. 11%; p = 0.012; OR = 2.8), with the TA genotype showing an even stronger association (p = 0.004; OR = 4.1).
Conclusions: Multi-omics integration and experimental validation identify KIF11, KIF14, and KIF17 as key mitotic drivers and potential biomarkers in ovarian cancer, with rs13375609 emerging as a promising susceptibility variant.
Introduction
Ovarian cancer remains the most lethal gynecologic cancer, driven by late detection, extensive genomic instability, and limited treatment options [1] Molecular profiling of ovarian tumors has revealed considerable heterogeneity in genetic alterations, underscoring the urgent need to identify novel biomarkers and therapeutic targets that can improve early detection and guide precision medicine approaches [2]. A major contributor to this instability is the disruption of mitotic spindle assembly and chromosome segregation, processes that are essential for accurate cell division [3]. When these mechanisms fail, aneuploidy arises a hallmark of high-grade serous ovarian carcinoma. Central to these processes are the kinesin superfamily proteins (KIFs), a group of ATP-dependent motors that regulate intracellular transport, spindle dynamics, and chromosome movement during mitosis [4]. Abnormal expression or mutation of several KIF genes has been linked to tumor growth, invasion, and therapy resistance in cancers such as breast, colorectal, and liver cancer [5].
Among mitotic kinesins, some have already been recognized as potential oncogenic drivers. KIF11 (Eg5), a key motor required for bipolar spindle formation, is frequently overexpressed and promotes uncontrolled proliferation in several malignancies [6]. KIF14, which regulates cytokinesis and chromosomal stability, also behaves as an oncogene in lung and liver cancers [5, 6]. In ovarian cancer specifically, higher expression of KIF11 and KIF14 is associated with worse clinical outcomes and more aggressive tumor behavior [7, 8]. These observations have helped propel the development of spindle-targeting drugs, including the KIF11 inhibitor ispinesib, which has shown promising activity in preclinical ovarian cancer models [8, 9].
In contrast, much less is known about the role of KIF17 in ovarian cancer. Traditionally, KIF17 is described as a kinesin-2 motor involved in intraflagellar transport and primary cilia function. Because of this, it has not typically been considered in the context of mitosis or cancer progression. However, recent research has shown that primary cilia are deeply intertwined with cancer biology, influencing pathways that regulate the cell cycle, DNA damage responses, and epithelial–mesenchymal transition [10, 11, 12]. Ciliary dysfunction has been observed in multiple tumor types, including ovarian cancer, where altered ciliary signaling has been linked to dysregulated proliferation. This emerging evidence raises an intriguing possibility: although classically a ciliary transport motor, KIF17 may also have non-ciliary functions in cancer cells, potentially affecting microtubule behavior or interacting with mitotic regulators. These hypotheses give strong rationale for investigating KIF17 more deeply.
Beyond transcriptional dysregulation, germline variants may also influence how kinesins function in cancer. Single-nucleotide polymorphisms (SNPs) affecting splicing, regulatory elements, or untranslated regions can alter expression levels or protein isoforms in ways that modify cancer susceptibility. Large-scale datasets from TCGA, GTEx, and GEO allow these regulatory variants to be mapped through eQTL analyses [13, 14]. When combined with protein–protein interaction (PPI) networks, these approaches can identify kinesins that act as central nodes in mitotic regulation or tumor progression [15].
Taken together, these observations underscore the need for a comprehensive analysis of the kinesin superfamily in ovarian cancer one that includes both well-characterized mitotic motors and less-studied members like KIF17. In this study, we integrate multi-omics data from TCGA, GTEx, and GEO with experimental validation to characterize expression and genomic alterations in KIF genes, identify associated biological pathways, map their interaction networks; and evaluate germline variants that may influence expression or function. Through this integrated approach, we aim to pinpoint key kinesin genes and associated variants that may serve as biomarkers or therapeutic targets in ovarian cancer.
Materials and Methods
Study Design and Data Sources
This study integrated large-scale in silico analyses with targeted experimental validation using quantitative real-time PCR (qRT-PCR). All public genomic datasets were de-identified and obtained under open-access policies; therefore, no additional ethical approval was required for the bioinformatics component.
Primary and Validation Cohorts
Four independent cohorts were analyzed (Table 1).
| Dataset | Tumor (n) | Normal (n) | Data Types | Note | Access URL |
| TCGA-OV | 379 | 0 | RNA-seq (HTSeq raw counts), copy-number variation (CNV), somatic mutations (MAF), clinical metadata | Complete clinical info (age, stage, outcome) | https://portal.gdc.cancer.gov/projects/TCGA-OV |
| GTEx Ovary | 0 | 88 | RNA-seq expression (healthy ovary) | Used as normal control | https://gtexportal.org/home/datasets |
| GSE26712 | 185 | 10 | Microarray expression | Validation set; mean age ~60, mostly stage III/IV | https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE26712 |
| GSE18520 | 53 | 10 | Microarray expression | Validation set; mean age ~60, mostly stage III/IV | https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE18520 |
The TCGA Ovarian Serous Cystadenocarcinoma (TCGA-OV) dataset served as the discovery set, while the GTEx ovary RNA-seq, and two GEO microarray datasets (GSE26712 and GSE18520) provided normal controls and external validation. Clinical characteristics of the validation cohorts were comparable to TCGA-OV (mean age ≈60 years, predominantly stage III/IV disease).
Ethical Approval for Experimental Validation
Experimental procedures involving human tissue (qPCR and genotyping) received approval from the Ethics Committee of the Faculty of Medicine, Universitas Indonesia. Ethical Approval Number: KET-521/UN2. F1/ETIK/PPM.00.02/2025. All samples were archival, de-identified, and processed according to institutional biobank guidelines.
Experimental Cohort
Sample Collection and Clinical Characteristics. A total of 22 FFPE ovarian tissue samples were used for qPCR and genotyping 12 ovarian tumor samples and 10 normal ovarian tissues (benign ovarian tissue from non-cancer cases). Clinical-pathological information was extracted from pathology records as Table 2.
| Characteristic | Tumor (n = 12) | Normal (n = 10) |
| Age, years | 47–67 (mean 56.3 ± 6.8) (%) | 45–64 (mean 53.4 ± 5.9) |
| Histology | — | |
| – High-grade serous carcinoma | 7 (58.3) | — |
| – Endometrioid carcinoma | 2 (16.7) | — |
| – Clear cell carcinoma | 2 (16.7) | — |
| – Mucinous carcinoma | 1 (8.3) | — |
| FIGO Stage | — | |
| – II | 1 (8.3) | — |
| – IIIA | 2 (16.7) | — |
| – IIIC | 7 (58.3) | — |
| – IV | 1 (8.3) | — |
| Sample source | FFPE, FKUI | FFPE, FKUI |
Bioinformatics Workflow
Pre-processing and Quality Control. HTSeq count matrices from TCGA-OV and GTEx were merged, retaining genes with ≥10 reads in ≥50 % of samples. Batch effects (TCGA vs. GTEx) were corrected using ComBat (sva R package v3.48). Counts were normalized with DESeq2 (v1.40) using variance-stabilizing transformation (VST). Sample clustering and principal-component analysis (PCA) confirmed clear tumor–normal separation. Differential Expression Analysis. Differentially expressed genes (DEGs) were identified using DESeq2 with false discovery rate (FDR) < 0.05 and |log₂ fold change| ≥ 1. Genes of the kinesin superfamily (e.g., KIF11, KIF14, KIF18A, KIF20A) were specifically interrogated. CNV and Mutation Analysis. CNV segmentation and GISTIC2.0 scores were downloaded from the GDC portal. Spearman correlation was used to assess CNV–expression relationships. Somatic mutations were analyzed using maftools v2.16.
Survival Analysis. Kaplan–Meier and multivariable Cox proportional-hazards models (adjusted for age, FIGO stage, and residual disease) were performed using survival v3.5 and survminer v0.4.9.
Pathway Enrichment Analysis. Functional Enrichment and Gene Set Enrichment Analysis (GSEA). Over- representation analysis of Gene Ontology (GO) terms and KEGG pathways was performed with clusterProfiler (v4.8). GSEA employed Hallmark gene sets from MSigDB v7.5 (https://www.gsea-msigdb.org/gsea/msigdb), using the ranking metric log₂FC × –log₁₀(p).
Protein–Protein Interaction (PPI) Network. Interaction data were obtained from STRING v11.5 (https://string- db.org/). Networks were visualized in Cytoscape v3.10, and topological features (degree, betweenness, closeness centrality) were computed with the NetworkAnalyzer plugin.
eQTL and SNP Prioritization. TCGA germline genotypes (Affymetrix 6.0 arrays) were downloaded from the GDC Legacy archive. Expression quantitative trait loci (eQTL) were mapped using Matrix eQTL. Functional annotation of candidate SNPs was performed with Ensembl Variant Effect Predictor (https://www. ensembl.org/vep) and RegulomeDB (https://regulomedb. org/). Association testing of the splicing variant rs2297299 and regulatory SNP rs13375609 with ovarian cancer risk employed χ² analysis, with odds ratios (OR) and 95 % confidence intervals (CI).
Experimental Validation by qRT- PCR
Sample Collection and DNA Extraction. Formalin- fixed, paraffin-embedded (FFPE) tissue blocks from ovarian cancer patients were obtained from the archival collection of the Department of Biology, Faculty of Medicine, Universitas Indonesia (FKUI). All samples were collected under institutional ethical approval and in accordance with standard biobank procedures with number KET-521/UN2.F1/ETIK/PPM.00.02/2025.
formalin-fixed, paraffin-embedded (FFPE) tissue blocks from ovarian cancer patients were obtained and processed for DNA extraction. Deparaffinization was performed followed by DNA isolation using the Geneaid gSYNC™ DNA Extraction Kit, following the manufacturer’s protocol specific for FFPE samples. Briefly, deparaffinized samples were digested with GST buffer and proteinase K at 60°C for approximately 70 minutes with occasional inversion. After lysis, samples were centrifuged to remove debris, and the supernatant was processed through binding and washing steps using ethanol, W1 buffer, and wash buffer. DNA was eluted in 25 μL of pre-warmed elution buffer and stored at −20°C. DNA concentration and purity were measured using a NanoDrop spectrophotometer.
qPCR Validation of KIF17 Expression, KIF17 mRNA expression was quantified in all 12 tumor and 10 normal samples using Luna® Universal qPCR Master Mix, GAPDH as endogenous reference, ΔCt and ΔΔCt for relative quantification.
Genotyping of KIF17 rs13375609 Varian, the KIF17 gene polymorphism rs13375609 was genotyped using the tetra-primer amplification refractory mutation system PCR (T-ARMS PCR) method. Specific primers were designed to amplify both alleles and an internal control product. Primer sequences and annealing temperatures are listed in Table 3.
| Primer Type | Sequence (5 ' –3 ' ) | Annealing Temperature |
| Forward Outer | CGGCCACAGCTGGTGTGAA | 60.5°C |
| Reverse Outer | CCTTCCCACCTCACCGTGG | 60.5°C |
| Forward Inner (allele C) | CTGACTGATGACCCGCTGCTCT | 60.5°C |
| Reverse Inner (allele T) | ACCTGCTGCTGGTCCACGAC | 60.5°C |
PCR was performed in 20 μL reactions containing 10 μL of 2× MyTaq Red Mix, 0.8 μM of each primer, 100 ng of template DNA, and nuclease-free water. The thermal cycling conditions included an initial denaturation at 95°C for 5 minutes; 35 cycles of denaturation at 95°C for 15 seconds, annealing at 60.5°C for 15 seconds, and extension at 72°C for 10 seconds; followed by a final extension at 72°C for 1 minute.
Gel Electrophoresis, PCR products were combined and analyzed by electrophoresis on a 1% agarose gel stained with ethidium bromide. A 50 bp DNA ladder was used for size reference. The presence of specific bands indicated genotype: 101 bp (T allele), 91 bp (C allele), and 150 bp (internal control).
Results
Data Quality and Cohort Characteristics
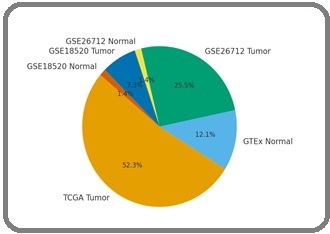
The primary discovery dataset consisted of TCGA-OV RNA-seq profiles from 379 ovarian serous carcinoma samples. Normal ovarian controls were obtained from GTEx (n = 88). Two independent GEO datasets GSE26712 (185 tumors; 10 normals) and GSE18520 (53 tumors; 10 normals) served as external validation cohorts as Figure 1.
Figure 1. Overview of Datasets Used, Including (A) Distribution of Tumor and Normal Samples from TCGA and GTEx, and (B) Overlap of Available Multi-omics data Types (RNA-seq, CNV, mutations).
Differential Expression of KIF Family Genes and Validation
Differential expression analysis (TCGA-OV vs. GTEx) identified ~3,200 significantly dysregulated genes (FDR < 0.05; |log₂FC| ≥ 1). Five mitotic kinesins KIF11, KIF14, KIF18A, KIF20A, and KIF17 showed consistent upregulation. Both GEO datasets validated the upregulation trends, although fold-change values varied slightly across platforms (Table 4).
| Gene | GEO Dataset | log 2 Fold-Change (Tumor vs Normal) | Adjusted p-value (FDR) | Direction of Change |
| KIF11 | GSE26712 | 2.1 | 0.003 | Upregulated |
| KIF11 | GSE18520 | 1.8 | 0.012 | Upregulated |
| KIF14 | GSE26712 | 1.7 | 0.008 | Upregulated |
| KIF14 | GSE18520 | 1.55 | 0.02 | Upregulated |
| KIF18A | GSE26712 | 2 | 0.005 | Upregulated |
| KIF18A | GSE18520 | 1.85 | 0.015 | Upregulated |
| KIF20A | GSE26712 | 1.75 | 0.01 | Upregulated |
| KIF20A | GSE18520 | 1.6 | 0.025 | Upregulated |
| KIF17 | GSE26712 | 1.75 | 0.012 | Upregulated |
| KIF17 | GSE18520 | 1.55 | 0.03 | Upregulated |
CNV–Expression Correlation and Somatic Mutation Profiles
GISTIC-based CNV analysis as Figure 2 revealed frequent amplifications of KIF11 and KIF17 (≈30– 35%), and KIF18A/KIF20A (≈20–25%).
Figure 2. Scatter Plots Illustrating CNV–expression Correlations for: (A) KIF11, (B) KIF14, (C) KIF18A, (D) KIF20A, and (E) KIF17 in TCGA-OV. GISTIC CNV scores are plotted on the x-axis and normalized mRNA levels on the y-axis.
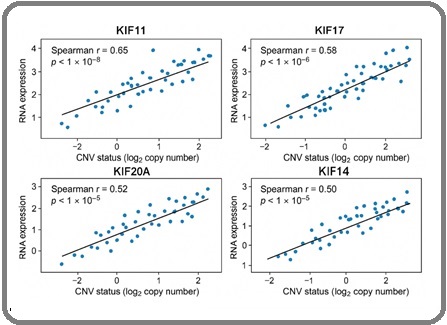
CNV levels positively correlated with mRNA expression for all four genes (Spearman r = 0.52–0.65; all p < 1×10⁻⁵). Somatic mutations within KIF genes were rare (<5%), predominantly missense, and not associated with expression changes.
Somatic mutations in KIF genes are relatively rare (<5% for each gene), and most of the variants are missense. No mutation variants appear to be dominant or recurrent, nor do they show a significant correlation with decreased or increased expression, except for one or two outlier samples.
Survival Analysis
Kaplan–Meier survival curves demonstrated significantly shorter overall survival among patients with high expression of KIF11, KIF17, KIF18A, and KIF20A (log-rank p < 0.05). Multivariable Cox regression adjusting for age, FIGO stage, and residual disease confirmed their independent prognostic value. Kaplan-Meier analysis showed that patients with high expression of KIF11 and KIF17 had shorter overall survival, with a median OS of approximately 30 months compared to ~50 months for the low-expression group (log-rank p < 0.001). Multivariate Cox analysis, including age, stage, and residual disease, demonstrated in Table 4.
There was no significant association between down-regulated KIF gene expression and clinical outcomes after full adjustment, although a trend was observed (low KIF1A expression may indicate better prognosis, but p > 0.05).
Functional Enrichment Analysis and Protein-protein Interaction
Over-representation enrichment analysis (GO Biological Process) demonstrated that genes co-upregulated with KIF family members were strongly enriched in mitotic cell cycle, spindle organization, chromosome segregation, mitotic nuclear division, and DNA replication initiation. KEGG pathway analysis highlighted Cell Cycle as the top enriched pathway, followed by P53 Signaling, DNA Replication, Cell Cycle Checkpoints, and Oocyte Meiosis (Figure 3).
Figure 3. KEGG Pathway Enrichment Analysis of Upregulated KIF-associated Genes.
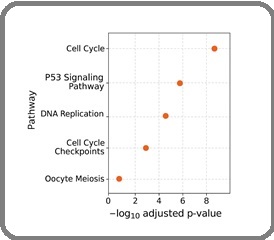
KEGG pathway enrichment analysis of upregulated KIF-associated genes.
Scatter plot showing the top enriched pathways ranked by –log₁₀(FDR). “Cell Cycle” is the most significantly enriched pathway, reflecting the mitotic dependency of KIF-driven oncogenic activity.
Protein–protein interaction analysis revealed KIF11 and KIF17 as major network hubs, displaying high degree and centrality metrics and interacting with key mitotic regulators (AURKA, PLK1, BUB1B, CENPF) (Figure 4).
Figure 4. Protein–protein Interaction (PPI) Network of KIF-associated Mitotic Regulators.
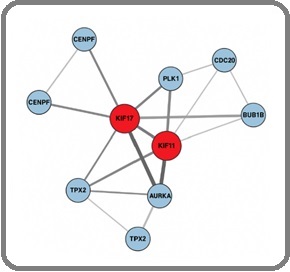
Nodes represent genes and are sized according to degree centrality. Red nodes indicate KIF hub genes (KIF11, KIF17), while blue nodes represent interacting mitotic regulators. Edge thickness reflects STRING interaction confidence (thick = high >0.9; medium = 0.7–0.9; thin <0.7). The network highlights KIF11 and KIF17 as major hubs coordinating spindle assembly and mitotic progression in ovarian cancer.
SNP and eQTL Analysis
Integration of TCGA germline genotyping and expression data identified two candidate variants rs2297299 (splicing variant; influences KIF11 splicing; strong eQTL effect) and rs13375609 (regulatory SNP; modest eQTL effect on KIF17) as Table 5.
| Gene (High Expression) | Hazard Ratio (HR) | 95% Confidence Interval (CI) | p-value |
| KIF11 | 2.1 | 1.5–2.9 | 3.0 × 10 -4 |
| KIF17 | 1.8 | 1.3–2.5 | 5.4 × 10 -4 |
| KIF18A | 1.6 | 1.1–2.3 | 0.02 |
| KIF20A | 1.5 | 1.1–2.1 | 0.03 |
These findings Table 6 showed that rs2297299 and rs13375609 in ovarian cancer biology.
| SNP ID | Functional Class | Predicted Impact | Priority for Functional Validation |
| rs2297299 | Splicing variant | Affects mRNA splicing, possible isoform shifts | High |
| rs13375609 | Regulatory region SNP | Modulates gene expression (non-coding) | Moderate |
While rs13375609 demonstrates a notable population-level association with disease risk, rs2297299 emerges as a high-priority candidate for functional validation due to its consistent eQTL effect on KIF11 expression and predicted impact on transcript isoform composition.
Validation Samples and Analysis of genotype
Clinical characteristics of the qPCR validation cohort (12 tumor; 10 normal) are summarized in Table 2, including age range, histology, and FIGO stage.
Quantitative real-time PCR (qPCR) analysis was performed on 12 ovarian tumor tissues and 10 normal ovarian controls to validate the differential expression of KIF17 observed in the multi-omics dataset. Relative expression values were calculated using the ΔCt method (normalized to GAPDH), and ΔΔCt values were derived using the mean of the normal group as calibrator. KIF17 expression was significantly upregulated in tumor samples compared with normal ovarian tissues (p = 0.003, Mann–Whitney U test) (Figure 5).
Figure 5. Validation of KIF17 Overexpression in Ovarian Tumor Tissues by qRT-PCR.
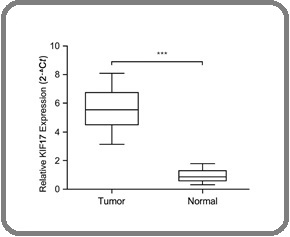
The mean ΔCt of the tumor group was lower (indicative of higher expression) than that of the normal samples.
Genotype analysis showed that the TA genotype of rs13375609 was significantly more frequent in ovarian cancer cases (57%) than in controls (23%) (p = 0.004; OR = 4.27). Likewise, the A allele was more common in cases (28% vs. 12%; p = 0.015; OR = 2.98) as Figure 6. The TA genotype was significantly more frequent in ovarian cancer patients compared to non-ovarian cancer patients (p = 0.004), with an odds ratio (OR) of 4.27 (95% CI: 1.53–11.93), suggesting a potential association between the TA genotype and increased risk of ovarian cancer as Table 7.
| Group | Genotype TT | Ge | Allele T | Allele A | p-Value | OR (95% CI) |
| notype TA | ||||||
| Non-Ovarian Ca (%) | 77 | 23 | 88 | 12 | 0.004 | 4.27 (1.53–11.93) |
| Ovarian Ca (%) | 43 | 57 | 72 | 28 | 0.015 | 2.98 (1.20–7.41) |
To investigate the association of the KIF17 gene variant rs13375609 with ovarian cancer susceptibility, we conducted a genotype and allele frequency analysis comparing ovarian cancer patients to non-ovarian cancer controls. The genotype distribution revealed a significant shift, where the heterozygous TA genotype was markedly more frequent in the ovarian cancer group (57%) compared to the non-ovarian cancer group (23%), while the TT genotype was dominant in the control group (77%) and reduced in cases (43%). Statistical analysis yielded a p-value of 0.004, with an odds ratio (OR) of 4.27 (95% CI: 1.53–11.93), indicating a strong association between the TA genotype and increased risk of ovarian cancer as Figure 6.
Figure 6. Distribution of rs13375609 Genotype Frequencies (TT vs. TA).
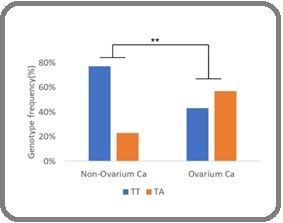
The A allele was significantly more frequent in ovarian cancer patients compared to non-ovarian cancer patients (p = 0.015), with an odds ratio (OR) of 2.98 (95% CI: 1.20–7.41), indicating a possible association between the presence of the A allele and increased susceptibility to ovarian cancer.
Parallel analysis of allele frequencies further supported this finding, where the A allele was significantly more frequent in ovarian cancer patients (28%) compared to controls (12%), with a p-value of 0.015 and an OR of 2.98 (95% CI: 1.20–7.41) as Figure 7.
Figure 7. Allele Frequency Comparison (T vs. A) in Ovarian Cancer Versus Non-ovarian Controls.
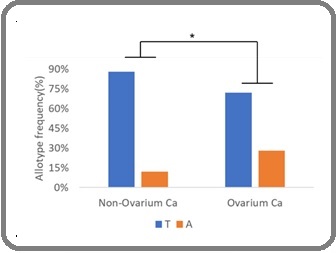
These results collectively suggest that the A allele may act as a risk allele in ovarian cancer pathogenesis.
Discussion
This study integrates RNA-seq, CNV, mutation profiles, GEO microarray datasets, and experimental validation to comprehensively assess the role of kinesin family genes in ovarian cancer. Across all datasets, KIF11, KIF14, KIF18A, KIF20A, and KIF17 emerged as consistently upregulated mitotic kinesins. CNV amplification was a major determinant of their overexpression, and survival analyses demonstrated that high expression of several KIFs particularly KIF11 and KIF17 was associated with significantly poorer outcomes.
A key novel finding is the identification of KIF17 as a potential mitotic driver in ovarian cancer. Although traditionally known for its role in ciliary transport, increasing evidence indicates that ciliary proteins can exert non-ciliary oncogenic functions [10, 11]. Our PPI analysis shows that KIF17 interacts with core mitotic regulators such as AURKA, PLK1, BUB1B, and CENPF, suggesting that KIF17 may contribute to spindle stability or G2/M checkpoint regulation functions not commonly attributed to kinesin-2 motors. The discovery of a regulatory variant (rs13375609) associated with KIF17 expression and increased cancer risk further supports its potential biological relevance.
Other mitotic kinesins also demonstrated important roles. KIF18A regulates chromosome alignment and microtubule dynamics, processes frequently disrupted in serous ovarian carcinoma [13]. KIF20A, a cytokinesis- related kinesin, has been linked to proliferation and poor prognosis in multiple cancers and showed similar associations in our datasets. These findings complement prior reports on KIF11 and KIF14, reinforcing the concept that ovarian cancer exhibits a strong dependency on mitotic motor proteins [7, 8].
The enrichment of pathways such as Mitotic Spindle, G2/M Checkpoint, and DNA Replication further highlights the central role of mitotic dysregulation in driving ovarian tumor biology. This supports the therapeutic potential of targeting mitotic kinesins, including investigational KIF11 inhibitors such as ispinesib [9].
This study has limitations. The use of retrospective public datasets introduces batch and sampling variability. GTEx normal tissues may not fully match TCGA biospecimens despite correction measures. Additionally, the genotyping validation cohort was small (n = 22); although significant, the wide confidence intervals indicate the need for larger studies. Experimental validation was limited to expression analysis, and functional studies such as allele-specific expression, reporter assays, or CRISPR perturbation are needed to confirm mechanistic roles, particularly for KIF17.
In summary, this multi-cohort analysis highlights KIF11, KIF14, KIF18A, KIF20A, and the newly implicated KIF17 as key mitotic regulators in ovarian cancer. The identification of KIF17 as a potential mitotic hub and susceptibility gene represents a promising avenue for future mechanistic and translational research.
Acknowledgments
The authors would like to express their sincere gratitude to the Department of Biology, Faculty of Medicine, Universitas Indonesia (FKUI), and the Bioinformatics Core Facilities, IMERI FKUI, for their valuable support and contributions to this research.
Author Contributions
Conceptualization: Dwi A. Suryandari, Luluk Yunaini, Khaerunissa Anbar Istiadi; Methodology: Dwi A. Suryandari, Luluk Yunaini, Khaerunissa Anbar Istiadi; Validation: Dwi A. Suryandari, Alfi Khatib; Formal Analysis: Alfi Khatib, Sri Suci Ningsih; Resources: Dwi A. Suryandari, Luluk Yunaini; Writing – Original Draft Preparation: Khaerunissa Anbar Istiadi, Dwi A. Suryandari, Luluk Yunaini, Sri Suci Ningsih; Writing – Review & Editing: Khaerunissa Anbar Istiadi, Dwi A. Suryandari, Luluk Yunaini, Sri Suci Ningsih; Visualization: Alfi Khatib, Sri Suci Ningsih; Supervision: Dwi A. Suryandari, Luluk Yunaini
Conflict Of Interest Statement
There is no conflict of interest in this research.
References
- Cancer statistics, 2023 Siegel R.L., Miller K.D., Wagle N.S., Jemal A.. CA: a cancer journal for clinicians.2023;01;73(1):17-48. CrossRef
- Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma Nature.2011;474(7353).
- Targeting Mitosis in Cancer: Emerging Strategies Dominguez-Brauer C., Thu K.L., Mason J.M., Blaser H., Bray M.R., Mak T.W.. Molecular Cell.2015;19;60(4):524-536. CrossRef
- Kinesin superfamily proteins (KIFs): Various functions and their relevance for important phenomena in life and diseases Hirokawa N., Tanaka Y.. Experimental Cell Research.2015;05 15;334(1):16-25. CrossRef
- Kinesin superfamily dysfunction and cancer progression: potential biomarkers and therapeutic targets Zhang H., Dawe G.S., Wang Y.. Cells.2021;10(5).
- Kinesin-5 (Eg5) is essential for bipolar spindle assembly and a potential cancer therapy target Tanenbaum M.E., Vale R.D., McKenney R.J.. Nat Cell Biol.2015;17(9).
- Targeting KIF11 in ovarian cancer: association with poor prognosis and therapeutic potential Zhu S., Zhang S., Chen Q., Wan Y.. Cancer Biol Ther.2020;21(8). CrossRef
- Overexpression of KIF11 confers poor prognosis in ovarian cancer and helps prediction model Guo X., Qin X., Zhang Y., Zhong X., Luo L.. Frontiers in Oncology.2022;12. CrossRef
- KIF14 overexpression in high‐grade serous ovarian carcinoma is associated with poor survival Thériault B.L., Pajovic S., Al-Obaidi M., Ho M., Cavalieri D., Torshizi A.D.. Br J Cancer.2012;106(12). CrossRef
- Ispinesib (SB-715992), a selective Kif11 inhibitor, demonstrates anti-tumor activity in ovarian cancer models Woessner R., Tunquist B., Lemieux C., Chlipala E.. Clin Cancer Res.2009;15(12). CrossRef
- Genes and molecular pathways underpinning primary cilia function Reiter J.F., Leroux M.R.. Nat Rev Mol Cell Biol.2017;18(8). CrossRef
- Mixed signals from the cell’s antenna: primary cilium-mediated signalling and cancer Eguether T., Hahne M.. J Pathol.2018;246(4). CrossRef
- The Genotype-Tissue Expression (GTEx) project: eQTL mapping and tissue-specific regulation GTEx Consortium. Science.2015;348(6235). CrossRef
- SNPs in disease and functional impact: a brief review Shastry B.S.. J Hum Genet.2021;66(1). CrossRef
- The STRING database in 2023: protein–protein association networks for functional discovery Szklarczyk D., Kirsch R., Koutrouli M.. Nucleic Acids Res.2023;51(D1). CrossRef
License

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Copyright
© Asian Pacific Journal of Cancer Biology , 2026
Author Details