Functional Impact of Interleukin -33 Polymorphism on IRF8 and TGF-β Regulation in Leukemia Patients
Download
Abstract
Introduction: The immune system plays an important role in controlling cancer development. Leukemia, a malignant disorder of hematopoietic tissues, is characterized by the abnormal proliferation of white blood cells. Unlike solid tumors, leukemia involves a systemic increase in aberrant blood cells rather than localized masses. This inflammatory condition is driven by immune mechanisms involving pathogenic cytokines, among which Interleukin-33 (IL-33) plays a complex and dual role. IL-33 influences the immune system and the tumor microenvironment (TME), potentially promoting or inhibiting leukemia progression depending on the context. A deeper understanding of its mechanisms could pave the way for innovative treatments targeting the IL-33 axis.
Methods: This study aimed to evaluate specific biochemical, hematological, and molecular markers in patients with leukemia. Conducted from June 1, 2024, to September 1, 2024, the study included 60 Iraqi patients with leukemia and 30 healthy controls. Serum levels of interferon regulatory factor 8 (IRF8) and transforming growth factor-beta (TGF-β) were measured using Enzyme-Linked Immunosorbent Assay (ELISA). Single-nucleotide polymorphisms (SNPs) of the IL-33 gene (rs928413) were assessed using Polymerase Chain Reaction (PCR) with a resistance mutation system.
Results: Revealed a highly significant increase in IRF8 levels in leukemia patients compared to controls (P < 0.0001), suggesting a strong association with the pathological state. While overall TGF-β levels did not differ significantly between groups, analysis of genetic polymorphisms indicated a potential influence of the IL-33 rs928413 genotype on TGF-β regulation in patients. Specifically, the GG genotype was associated with the highest IRF8 and TGF-β levels in patients. Although allele distribution of IL-33 rs928413 did not show a direct association with increased risk in this sample, the findings suggest a complex interplay between IL-33 gene polymorphisms, IRF8, and TGF-β in leukemia.
Conclusion: Leukemia patients exhibit elevated immune markers, particularly IRF8, which may play a significant role in disease onset and progression. Further studies are warranted to elucidate the molecular mechanisms linking IL-33 gene polymorphisms to IRF8 and TGF-β function in leukemia, and to explore their potential as therapeutic targets or prognostic markers.
1. Introduction
Acute myeloid leukemia (AML) is characterised by several molecular and cytogenetic abnormalities. Cytokines and growth factors produced in the medullary microenvironment govern cell survival, proliferation, and differentiation; hence, polymorphisms in their associated genes may affect cancer risk [1]. AML is an aggressive haematological malignancy characterised by a death rate of 60% and a recurrence rate of 30-40% among patients in remission [2]. Genetic determinants are pivotal in illness prognosis, rendering genetic studies indispensable for disease evaluation and management [2]. AML is the most prevalent form of acute leukemia in adults, characterised by rapid progression and, if left untreated, would result in the patient’s demise within weeks or months [3].
Characterization of critical transcription factors in AML may identify potential alternative targets to modulate specific gene-regulation pathways/inhibitors for AML. These drugs are expected to improve overall survival in AML patients [3]. IRF8, identified as a member of the interferon IRFs transcription factor family, exerts various functions in hematopoiesis and participates in type I interferon response [4].
IRF8 is primarily recognised for its role in the formation and maturation of myeloid cells [5]. It modulates lineage- committed progenitors to facilitate monocyte maturation and inhibit neutrophil generation [6, 7]. IRF8 has been previously characterised as a tumour suppressor across many cancer types [8, 9], including chronic myeloid leukemia (CML) and acute promyelocytic leukemia [10, 11], such that disruption of IRF8 activity promotes leukemogenesis [12].
TGFB is a key modulator of normal hemopoiesis. An aberrant TGFB pathway has been implicated in the pathogenesis of various hematological malignancies, including myelofibrosis, acute myeloid leukemia, and lymphoid disorders. TGF-β, which has long been considered an inhibitor of proliferation in hematopoiesis, also induces differentiation and apoptosis to maintain homeostasis. Tumors often acquire intrinsic resistance to homeostatic TGF-β signaling, which counteracts its tumor-suppressive actions [13].
In addition, elevated TGF-β levels contribute to cancer progression by modulating the immune response and altering the tumour microenvironment [14]. Surface receptors of TGF-β are down-modulated, SMAD signaling proteins are abnormal, and the CDKN2B gene is deleted or silenced at both the genetic and epigenetic levels in the context of malignant B-cell subtypes. The TGF-β in the microenvironment provides a protumoral niche via trans differentiation of stroma, natural killer (NK) cells, and T cells. There is mounting evidence for extensive mutual communication between microenvironmental cells and tumorigenic B cells [15]. Revealing intracellular communication and context-dependent TGFβ signaling could help elucidate disease pathogenesis, thereby offering a new potential therapeutic strategy.
IL-33, a member of the IL-1 family of cytokines, is critically involved in regulating immunity and inflammation [16]. Its receptor, suppression of tumorigenicity 2 (ST2), a member of the interleukin-1 receptor family, transduces signals through the MyD88 pathway, thereby activating downstream inflammatory cascades [17]. IL-33 has recently been investigated in several cancers, including leukemia [18]. IL-33 is a key regulator of the immune TME in leukemia and modulates the activity of immune cells such as regulatory T cells (Tregs), myeloid-derived suppressor cells (MDSCs), and natural killer cells [18, 19]. The impact of IL-33 on leukemia is either promoting or inhibiting, depending on the precise context [5]. Importantly, IL-33 contributes to leukemia cell survival by enhancing the suppressive activity of Tregs and MDSCs, thereby inhibiting anti-tumour immunity [18]. Despite the potential immune cell-activating functions of IL-33, e.g., in T cells, NK cells, and macrophages, which may aid anti-leukemic responses, it can also promote leukemia in a setting of chronic inflammation. This is believed to be due to IL-33-induced production of tumour-promoting cytokines that enhance the survival and proliferation of cancer cells [20, 21]. The purpose of this study was to assess different haematological, biochemical and molecular indices in leukemia patients. The case-control study was conducted between March and April 2024, using blood samples from 60 patients with leukemia and 30 controls to analyses serum IL-33 levels and SNPs. Serum concentrations of IRF8 and TGF-β were measured by ELISA to determine whether they were associated with the patients. Meanwhile, PCR technology using a resistance mutation system was used to amplify the primer set for SNP detection.
2. Materials and Methods
2.1 Sample Collection
A total of 90 samples were collected from individuals between June 1, 2024, and September 1, 2024. Participants’ ages ranged from 16 to 45 years. The samples were divided into two groups: 60 samples from leukemia patients who visited a specialized leukemia hospital in 2024 and had their diagnoses confirmed by specialists after appropriate chemical analyses, and 30 samples from healthy individuals (control group).
2.2 Serum Preparation
Five milliliters (5 mL) of venous blood were drawn from each individual (whether a leukemia patient or a healthy individual). These samples were divided into two parts for analysis. First, 2 mL of venous blood was placed in test tubes containing EDTA anticoagulant and stored for subsequent molecular study. In the second part, 3 mL of blood was placed in test tubes containing silicone gel-tubes. Subsequently, the samples were centrifuged at 3500 rpm to obtain serum. The serum was then transferred to Eppendorf test tubes and stored at -20°C. All relevant sample information was recorded and documented in preparation for biochemical assays. In this study, the concentrations of IRF8 and TGF-β were measured by ELISA using a range of ready-made solutions following the instructions provided for each assay.
2.2.1 Reagents
The serum levels of IRF8 and TGF-β were determined using commercially available ELISA kits according to the manufacturers’ instructions. The IRF8 ELISA kit was purchased from Sigma-Aldrich, St. Louis, MO, USA), while the TGF-β ELISA kit was obtained from R&D Systems (Minneapolis, MN, USA).
2.2.2 Ethics statement
Study approval was granted by the Research Ethics Committee of the Iraqi Ministries of Environment, Health, Higher Education, and Scientific Research; prior approval was obtained from the University of Tikrit Board; and the study adhered to the Declaration of Helsinki’s ethical standards for human testing, as required by the Health Insurance Portability and Accountability Act. The Data Safety Monitoring Board conducted routine reviews, including sample collection by NIH criteria and with consent from the University of Tikrit Board.
2.3 Molecular Study
2.3.1 DNA Extraction and Genotyping of the SNP rs928413 in the IL-33 Gene
Genomic DNA was isolated from peripheral blood using a Geneaid microarray kit (Geneaid, Taiwan). The rs928413 fragment of the IL-33 gene was amplified using two sets of primers in an allele-specific polymerase chain reaction (PCR). The primer sequences used were: IF: CAGTAGTTAGCTACTTTTTAATAGTTCCG, IR: CTGCCTTGGCCA AT GCTATT, OF: TGAATAGCATGGAGTAACTTGTGG, and OR: GCAAGTGTTTTGAAGAGGAAACTC, resulting in a resulting amplification size of 350 base pairs for the outlier, 226 base pairs for the GG genotype, and 173 base pairs for the AA genotype.
2.3.2 Polymerase Chain Reaction (PCR) Protocol
PCR was performed in a total reaction volume of 25 µL using a commercially prepared master mix containing Taq DNA Polymerase and essential reaction components. To this mix, 1 µL of each primer (at a concentration of 10 pmol/µL) and 2 µL of template DNA (concentration: 50 ng/µL) were added. The final volume was adjusted using DNase/RNase-free water. Amplification was carried out using a thermal cycler under the following program: initial denaturation at 94°C for 5 minutes, followed by 35 cycles of: denaturation at 94°C for 30 seconds, primer annealing at 57°C for 30 seconds, extension at 72°C for 45 seconds, and a final extension at 72°C for 5 minutes. Post-amplification, PCR products were analyzed by agarose gel electrophoresis using a 1.5% agarose gel stained with RedSafe DNA. Gel imaging was performed using a gel documentation system.
2.3.3 Gel electrophoresis
The reaction yielded bands of 350 bp for the outer fragment, 226 bp for the G allele, and 173 bp for the A allele. The presence of both allele-specific bands (226 bp and 173 bp) indicated a heterozygous GA genotype, while the presence of a single band corresponded to a homozygous genotype for one of the alleles.
2.4 Statistical analysis
The sample groups were found to demonstrate Hardy– Weinberg equilibrium for the rs928413 polymorphism using the chi-square test. Quantitative results are indicated as mean ± standard deviation. GraphPad Prism Software version 10.3 (GraphPad Software, USA) was used for the statistical analysis of the clinical and biochemical data obtained. The student’s t-test was used and the difference considered significant at P< 0.05. The rs3741664 variant’s genotypes and allele frequencies were calculated as percentages. To analyze the differences between controls and patients (confidence interval), the odds ratio (OR) and its 95% confidence interval (CI) were utilized.
3. Results and Discussion
3.1 Results
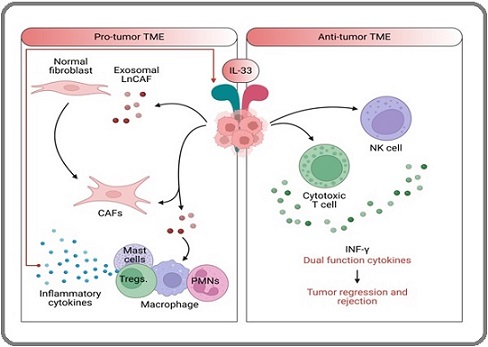
This study aimed to evaluate IRF8 and TGF-β levels in leukemia patients compared with healthy individuals and to analyze the single-nucleotide polymorphism rs928413 in the IL-33 gene. The biological role of IL-33 in the tumor microenvironment is illustrated in Figure 1.
Figure 1. This Illustration Depicts the Bifunctional Function of IL-33 in the Tumor Microenvironment (TME). On the left side of a pro-tumor TME, normal fibroblasts get activated into cancer-associated fibroblasts (CAFs) that further interact with mast cells, regulatory T cells (Tregs), and polymorphonuclear leukocytes (PMNs). This communication, combined with exosomal IL-33 released by CAFs, leads to the secretion of proinflammatory cytokines by CSCs, thereby favoring tumor growth. In addition, IL-33 supports this pro-tumor process. On the right, in an anti-tumor TME, IL-33 stimulates both natural killer cells (NK) and cytotoxic T cells. These cells produce interferon-gamma (IFN-γ), a cytokine with dual functions that can lead to both tumor regression and rejection. This shows that the TME context determines whether IL-33 acts as a promoter or a suppressor of tumor growth..
The results revealed clear differences between the patient and healthy control groups, highlighting the potential roles of these biomarkers in leukemia progression. Our findings demonstrated that the mean IRF8 levels in the patient group were 41.10 ± 1.33, significantly higher than the mean of 29.27 ± 2.06 observed in the control group (P < 0.0001). This highly statistically significant difference indicates a strong association between elevated IRF8 levels and the pathological state in patient samples. Furthermore, the range of values in patients (19.74 to 87.23) was broader than in healthy samples (2.851 to 47.16), suggesting greater variability in measured protein levels among patients, which may be attributed to differences in gene expression Table 1 and Figure 2.
| IRF 8 | Control | Patients | IL 33 | ||
| Number of values | 30 | 60 | GG | AG | AA |
| Minimum | 2.851 | 19.74 | 5 | 51 | 4 |
| Maximum | 47.16 | 87.23 | 38.22 | 19.74 | 33.34 |
| Range | 44.31 | 67.49 | 54.64 | 87.23 | 46.65 |
| Mean | 29.27 | 41.1 | 16.42 | 67.49 | 13.3 |
| Std. Deviation | 11.32 | 10.3 | 44.85 | 41.01 | 37.58 |
| Std. Error of Mean | 2.067 | 1.33 | 7.433 | 10.77 | 6.255 |
| P value | <0.0001 |
Figure 2. Genotypes Generated by T-ARMS PCR on Electrophoresis 1.5% Agarose Gel for the IL33 Gene (rs928413) Accompanied by a 100 Base pair DNA Ladder Marker.
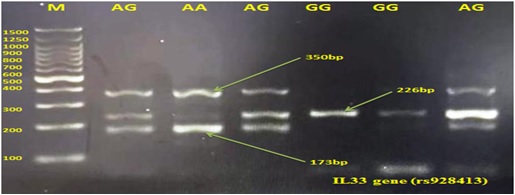

Table 2 shows the relationship between IRF8 and the rs928413 gene polymorphism in patients, showing that the GG genotype had the highest mean IRF8 levels (44.85 ± 3.32), followed by the AG genotype (41.01 ± 1.51), while the AA genotype had the lowest (37.58 ± 3.13). This gradient in the means suggests a possible influence of the gene mutation on IRF8 expression Figure 1.
| IRF 8, TGF-β Control | AA | AG |
| Number of values | 2 | 28 |
| Minimum | 30.81 | 2.851 |
| Maximum | 32 | 47.16 |
| Range | 1.191 | 44.31 |
| Mean | 31.41 | 29.12 |
| Std. Deviation | 0.842 | 11.72 |
| Std. Error of Mean | 0.5954 | 2.214 |
| P value | 0.7877 |
In the healthy control group, the mean IRF8 and TGF-β in AA carriers was 31.41 ± 0.59 compared to 29.12 ± 2.21 in AG carriers. No individuals with the GG genotype were found in the control group. However, the difference was not statistically significant (P = 0.7877), suggesting that the mutation’s effect in healthy individuals might be limited or undetectable.
Regarding TGF-β, our results showed no statistically significant difference between patients (44.71 ± 1.63) and healthy controls (42.69 ± 1.73; P = 0.4431). This suggests that the levels of this factor are not significantly associated with leukemia in the studied samples as shown in Table 3.
| TGF-β | Patients | Control | AA | AG | GG |
| Number of values | 60 | 30 | 4 | 51 | 5 |
| Minimum | 19.27 | 25.24 | 19.27 | 24.95 | 48.14 |
| Maximum | 69.28 | 59.7 | 35.17 | 69.23 | 69.28 |
| Range | 50.01 | 34.46 | 15.9 | 44.27 | 21.14 |
| Mean | 44.71 | 42.69 | 29.15 | 44.49 | 59.32 |
| Std. Deviation | 12.69 | 9.484 | 7.138 | 11.79 | 9.476 |
| Std. Error of Mean | 1.638 | 1.732 | 3.569 | 1.65 | 4.238 |
| P VALUE | 0.4431 |
The relationship between TGF-β and the genetic polymorphism results in patients showed that the GG genotype had the highest mean of 59.32 ± 4.23, followed by the AG genotype with 44.49 ± 1.65, and the AA genotype with 29.15 ± 3.57. These findings support the hypothesis that the gene mutation may influence TGF-β regulation in patients, as shown in Table 3.
The distribution of A and G alleles was similar between patients and healthy individuals, with no statistically significant difference (P = 0.6261), suggesting that the allele distribution is not directly associated with increased risk in this sample. These results suggest an association between IRF8 levels and the rs928413 gene polymorphism in patients, suggesting that this genetic polymorphism may also affect TGF-β regulation. However, the relationship with allele distribution remains unclear and requires expanded studies for confirmation, as shown in Table 4.
| Alleles | Study participants | |||||||||||||||
| Patients NO. | Allele frequency | Healthy control No. | Allele frequency | Allele frequency in total study participants | OR | 95% CI | χ2 | p-value | ||||||||
| Lower | Upper | |||||||||||||||
| A (reference) | 59 | 0.49 | 32 | 0.53 | 0.5 | 1 | ||||||||||
| G | 61 | 0.51 | 28 | 0.47 | 0.5 | 0.8493 | 0.4507 | 1.664 | 0.2374 | 0.6261 | ||||||
| Total | 120 | 1 | 60 | 1 | 1 | |||||||||||
| Genotype | Study participants | |||||||||||||||
| Patients | Healthy control NO. | OR | 95% CI | χ2 | p-value | |||||||||||
| No. | Lower | Upper | ||||||||||||||
| AA (reference) | 4 | 2 | 1 | |||||||||||||
| AG | 51 | 28 | 0.9107 | 0.1653 | 4.12 | 0.01087 | 0.917 | |||||||||
| GG | 5 | 0 | 0 | 0 | 2.469 | 2.037 | 0.1535 |
Upon studying the gene polymorphism and its relationship with TGF-β levels, it was observed that the GG genotype was absent in the healthy group but was present in patients at a significant level. However, the differences did not reach statistical significance (P = 0.1535). Similarly, there was no significant difference (Figure 3 and 4).
Figure 3. Serum AFR8 and TGF Levels among Patients and Control and Its Correlation to IL33 Gene (rs928413) Polymorphism.
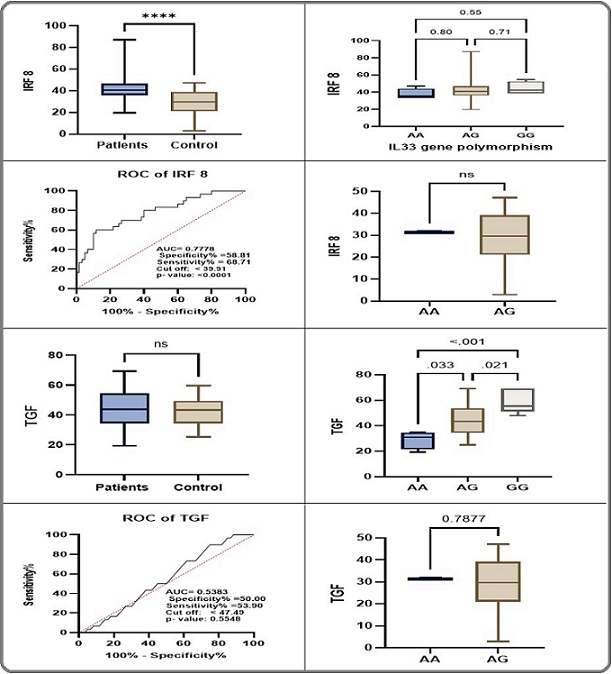
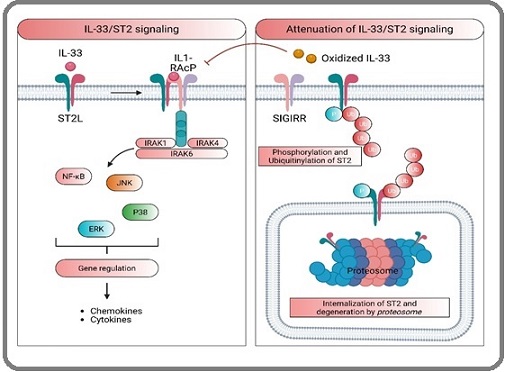
Figure 4. Activation of IL-33/ST2 Signaling Begins when IL-33 binds to ST2L, Triggering a Conformational Change that Recruits IL-1RacP to form a Heterodimer. This complex recruit’s adaptor molecules like MyD88, IRAK1, IRAK4, and TRAF6. The subsequent activation of transcription factors such as NF-κB, JNK, ERK, and p38 leads to the expression of genes for various cytokines, chemokines, and growth factors. The pathway can be attenuated in several ways: the molecule SIGIRR can disrupt the ST2L/IL1RAcP heterodimer; phosphorylated ST2L is rapidly internalized and degraded by the proteasome with the help of the E3 ligase FBXL19; and extracellular IL-33 can be sequestered by sST2 acting as a decoy or oxidized at its cysteine residues, preventing its binding to the ST2L receptor.
3.2 Discussion
Our results are consistent with those of Hartung et al. (2024), who found that IRF8 is essential for hematopoietic cell differentiation and for preventing myeloid leukemia [22]. Liss et al. (2021) revealed that IRF8 may function as a novel diagnostic biomarker and therapeutic target for certain types of AML [3]. The transcription factor IRF8 regulates AML cell sensitivity to LSD1 inhibition, a process implicated in the differentiation induced by all-trans retinoic acid (ATRA) and in the response to treatment [22]. Furthermore, IRF8 ablation suppressed AML cell proliferation, indicating that IRF8 is a tumor suppressor [23]. IRF8 is a well-established monoblastic marker in acute monocytic leukemias, underscoring its diagnostic utility [24]. Cumulatively, these data highlight the multifaceted role of IRF8 in leukemia pathogenesis and as a therapeutic target. The elevated expression of IRF8 we observed in leukemia patients suggests it is a compensatory mechanism or an immune response dysregulation, and this should be clarified by identifying which cell types and molecular methods are involved.
The polymorphism may affect IRF8 expression in leukaemia, but its effects on healthy individuals may be minor or necessitate specific environmental triggers to become apparent. Additional research into gene-environment interactions may yield greater elucidation. The varying effects of the rs928413 polymorphism on IRF8 levels in patients, especially the elevated levels associated with the GG genotype, suggest a possible genetic predisposition or modifying influence that may affect disease progression or therapeutic response. This warrants further functional studies to elucidate the molecular mechanisms by which this specific SNP affects IRF8 expression and activity.
Despite the absence of statistically significant differences in overall TGF-β levels between the two groups, recent research indicates that the TGF-β signaling pathway plays a dual role at different stages of cancer progression, potentially suppressing tumor formation in early stages and promoting its advancement in later stages [25]. A comprehensive characterization of TGFB1 across hematological malignancies revealed that TGFB1 is broadly dysregulated in these conditions and might regulate the immune microenvironment in a cancer-type- specific manner [26, 27]. Furthermore, TGFB1 levels are significantly elevated in treatment-resistant AML patients, and inhibition of the TGFB signaling pathway can enhance treatment efficacy, suggesting its critical role in drug resistance and therapeutic strategies [28]. These findings suggest that while basal levels may not differ significantly, the functional implications of TGF-β in leukemia, particularly in advanced or resistant forms, warrant further investigation. The lack of a significant difference in our study might be due to the heterogeneous nature of leukemia, where TGF-β\’s role could vary depending on the specific subtype or stage of the disease. Future studies could explore TGF-β levels in specific leukemia subtypes to uncover more nuanced associations.
IL-33, a constituent of the IL-1 cytokine family and a multifunctional cytokine, is pivotal in sustaining host homeostasis and in pathological states, including allergies, infectious diseases, and cancer, by influencing various immune cell types and enhancing type 1 and 2 immune responses. IL-33 is swiftly produced by both immune and non-immune cells in response to stress, functioning as an “alarmin” by binding to its receptor, suppression of tumorigenicity 2 (ST2), therefore initiating downstream signalling pathways and activating inflammatory and immunological responses. IL-33 is recognized for its dual immune regulatory functions in several disorders, exhibiting both pro- and anti-tumorigenic actions, that likely depend on its primary target cells, IL-33/sST2 expression levels, cellular context, and the cytokine microenvironment [29]. The dual role of IL-33 in cancer, as highlighted by recent research, where it can either promote or inhibit tumor progression depending on the context, further complicates the interpretation of these genetic associations [30]. Future studies should aim to elucidate the precise mechanisms by which IL-33 gene polymorphisms influence the expression and function of both IRF8 and TGF-β in leukemia, thereby enabling personalized therapeutic approaches [31]. The absence of the GG genotype in the healthy control group, coupled with its presence and association with higher TGF-β levels in patients, suggests a potential role for this genotype in leukemia pathogenesis, possibly by influencing the immune microenvironment. This observation warrants further investigation in larger cohorts to confirm its significance and explore its clinical implications.
In conclusion, leukemia patients exhibit elevated immune markers, particularly IRF8 and TGF-β, which may play a significant role in the onset and progression of the disease. Further studies are needed to evaluate their importance in individuals affected by leukemia. Future research should focus on understanding the molecular mechanisms linking IL-33 gene polymorphisms to IRF8 and TGF-β expression and function in the context of leukemia, and on exploring the potential of these markers as therapeutic targets or prognostic indicators.
Abbreviations
IL-33, Interleukin-33; IL-1, Interleukin-1; TME, Tumor microenvironment; IRF8, Interferon Regulatory Factor 8; TGF-β, Transforming Growth Factor-beta; ELISA, Enzyme-Linked Immunosorbent Assay; SNPs, Single Nucleotide Polymorphisms; PCR, Polymerase Chain Reaction; AML, Acute myeloid leukemia; CML, Chronic myeloid leukemia; NK cells, Natural killer cells; MDSCs, Myeloid-derived suppressor cells; CAFs, Cancer-associated fibroblasts; PMNs, Polymorphonuclear leukocytes; IFN- γ, Interferon-gamma; ST2, Suppression of tumorigenicity 2; ATRA, All-Trans Retinoic Acid.
Declarations
Ethics approval and consent to participate
Study approval was granted by the Research Ethics Committee of the Iraqi Ministries of Environment, Health, Higher Education, and Scientific Research; prior approval was obtained from the University of Tikrit Board; and the study adhered to the Declaration of Helsinki’s ethical standards for human testing, as required by the Health Insurance Portability and Accountability Act. The Data Safety Monitoring Board conducted routine reviews, including sample collection by NIH criteria and with consent from the University of Tikrit Board.
Consent for publication
Not applicable. This study does not include any individual person’s data in any form (including individual details, images, or videos).
Availability of data and material
The data that support the findings of this study are not publicly available due to [ethical/legal/privacy] restrictions but are available from the corresponding author on reasonable request.
Competing interests
The authors declare that they have no conflict of interests.
Funding
The author declare that no funds, grants, or other support were received during the preparation of this manuscript. The author have no relevant fnancial or non- fnancial interests to disclose.
References
- Interleukin-4 gene intron 3 VNTR polymorphism in adult acute myeloid leukemia Safwat NA , Najjar MRE , Saeed AM , Mohamed HS . Egyptian Journal of Medical Human Genetics.2022;23(1). CrossRef
- Evaluation of different in silico tools for the assessment of deleterious variants in acute myeloid leukemia Qureshi W, Irfan M, Khan IA , Shakeel M. Egyptian Journal of Medical Human Genetics.2025;26(1). CrossRef
- IRF8 Is an AML-Specific Susceptibility Factor That Regulates Signaling Pathways and Proliferation of AML Cells Liss F , Frech M , Wang Y , Giel G , Fischer S , Simon C , Weber LM , et al . Cancers.2021;13(4). CrossRef
- Type I Inteferon Gene Induction by the Interferon Regulatory Factor Family of Transcription Factors Honda K, Takaoka A, Taniguchi T. Immunity.2006;25(3). CrossRef
- Role of IRF8 in immune cells functions, protection against infections, and susceptibility to inflammatory diseases Salem S, Salem D, Gros P. Human Genetics.2020;139(6-7). CrossRef
- IRF-8 extinguishes neutrophil production and promotes dendritic cell lineage commitment in both myeloid and lymphoid mouse progenitors Becker AM , Michael DG , Satpathy AT , Sciammas R, Singh H, Bhattacharya D. Blood.2012;119(9). CrossRef
- IRF8 acts in lineage-committed rather than oligopotent progenitors to control neutrophil vs monocyte production Yáñez A, Ng MY , Hassanzadeh-Kiabi N, Goodridge HS . Blood.2015;125(9). CrossRef
- Epigenetic disruption of interferon-γ response through silencing the tumor suppressor interferon regulatory factor 8 in nasopharyngeal, esophageal and multiple other carcinomas Lee K Y, Geng H, Ng K M, Yu J, Van Hasselt A, Cao Y, Zeng Y-X, et al . Oncogene.2008;27(39). CrossRef
- The tumor suppressor interferon regulatory factor 8 inhibits β-catenin signaling in breast cancers, but is frequently silenced by promoter methylation Luo X, Xiong X, Shao Q, Xiang T, Li L, Yin X, Li X, Tao Q, Ren G. Oncotarget.2017;8(30). CrossRef
- Identification of IRF8 as a potent tumor suppressor in murine acute promyelocytic leukemia Gaillard C, Surianarayanan S, Bentley T, Warr MR , Fitch B, Geng H, Passegué E, De Thé H, Kogan SC . Blood Advances.2018;2(19). CrossRef
- All-trans retinoic acid and arsenic trioxide fail to derepress the monocytic differentiation driver Irf8 in acute promyelocytic leukemia cells Liu X, Chen J, Yu S, Yan L, Guo H, Dai J, Zhang W, Zhu J. Cell Death & Disease.2017;8(5). CrossRef
- The Prognostic Significance of IRF8 Transcripts in Adult Patients with Acute Myeloid Leukemia Pogosova-Agadjanyan EL , Kopecky KJ , Ostronoff F, Appelbaum FR , Godwin J, Lee H, List AF , et al . PLoS ONE.2013;8(8). CrossRef
- Transforming Growth Factor-Beta Orchestrates Tumour and Bystander Cells in B-Cell Non-Hodgkin Lymphoma Timmins MA , Ringshausen I. Cancers.2022;14(7). CrossRef
- Exploring TGF-β Signaling in Cancer Progression: Prospects and Therapeutic Strategies Sheikh K, Amjad M, Irfan M, Anjum S, Majeed T, Riaz M, Jassim A, et al . OncoTargets and Therapy.2025;Volume 18. CrossRef
- Transforming growth factor-β signaling: from tumor microenvironment to anticancer therapy Chan MK , Chan EL , Ji ZZ , Chan AS , Li C, Leung K, To K, Tang PM . Exploration of Targeted Anti-tumor Therapy.2023. CrossRef
- Interleukin‐33 (IL‐33): a nuclear cytokine from the IL‐1 family. Immunological reviews Cayrol C, Girard J. Immunological Reviews.2018;281(1). CrossRef
- IL-33 biology in cancer: An update and future perspectives Yeoh WJ , Vu VP , Krebs P. Cytokine.2022;157. CrossRef
- Dual targeting of tumoral cells and immune microenvironment by blocking the IL-33/IL1RL1 pathway Fu D, Jiang H, Long A, Harris E, Guo H, Capitano ML , Wrangle J, et al . Nature Communications.2025;16(1). CrossRef
- Interleukin-33 in Malignancies: Friends or Foes? Shen J, Liu J, Zhang G. Frontiers in Immunology.2018;9. CrossRef
- The Janus Face of IL-33 Signaling in Tumor Development and Immune Escape Choi M, Sosman JA , Zhang B. Cancers.2021;13(13). CrossRef
- Anti-Tumorigenic Activities of IL-33: A Mechanistic Insight Andreone S, Gambardella AR , Mancini J, Loffredo S, Marcella S, La Sorsa V, Varricchi G, Schiavoni G, Mattei F. Frontiers in Immunology.2020;11. CrossRef
- The Transcription Factor IRF8 Regulates the Sensitivity of AML Cells to LSD1 Inhibition and All-Trans Retinoic Acid Hartung EE , Singh K, Coopersmith S, Avrumutsoae J, Varatharajan S, Pishyar S, Prabagaran P, et al . Blood.2024;144(Supplement 1). CrossRef
- Loss of IRF8 inhibits the growth of acute myeloid leukemia cells Zhuang H, Li F, Xu Y, Pei R, Chen D, Liu X, Li S, et al . Annals of Hematology.2023;102(5). CrossRef
- IRF8 Is a Reliable Monoblast Marker for Acute Monocytic Leukemias, But Does Not Discriminate Between Monoblasts and Plasmacytoid Dendritic Cells Tzankov A , Facchetti F , Mühleisen B , Dirnhofer S . The American journal of surgical pathology.2022;46(5). CrossRef
- TGF-β Signaling in Cancer: Mechanisms of Progression and Therapeutic Targets Cecerska-Heryć E , Jerzyk A , Goszka M , Polikowska A , Rachwalska J , Serwin N , Wojciuk B , Dołęgowska B . International journal of molecular sciences.2025;26(15). CrossRef
- Comprehensive characterization of TGFB1 across hematological malignancies Wang C, Zhang Z, Zhang Y, Zheng L, Liu Y, Yan A, Zhang Y, et al . Scientific Reports.2023;13(1). CrossRef
- Role of sgk1 in cancer: a bibliometric analysis from 2013 to 2023—review article Abo- Elenien WI , Badawy SG , Abouelenin O, Hussein FK , Kumari S. Egyptian Journal of Medical Human Genetics.2024;25(1). CrossRef
- The roles and mechanisms of TGFB1 in acute myeloid leukemia chemoresistance Liang X, Zhou J, Li C, Wang H, Wan Y, Ling C, Pu L, et al . Cellular Signalling.2024;116. CrossRef
- Dual Immune Regulatory Roles of Interleukin-33 in Pathological Conditions Guo H, Bossila EA , Ma X, Zhao C, Zhao Y. Cells.2022;11(20). CrossRef
- Exogenous IL-33 overcomes T cell tolerance in murine acute myeloid leukemia Qin L, Dominguez D, Chen S, Fan J, Long A, Zhang M, Fang D, et al . Oncotarget.2016;7(38). CrossRef
- IL-33–ST2 signaling promotes stemness in subtypes of myeloid leukemia cells through the Wnt and Notch pathways Naef P, Radpour R, Jaeger-Ruckstuhl CA , Bodmer N, Baerlocher GM , Doehner H, Doehner K, et al . Science Signaling.2023;16(800). CrossRef
License

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Copyright
© Asian Pacific Journal of Cancer Biology , 2026
Author Details