Association between MICA rs2596542 and MTHFR rs1801133 Polymorphisms with the Risk of Hepatocellular Carcinoma
Download
Abstract
Background: Hepatocellular carcinoma (HCC) susceptibility is influenced by various risk factors, including viral infections, alcohol consumption, tobacco use, and genetic predisposition. The MICA rs2596542 and MTHFR rs1801133 gene polymorphisms have been implicated in HCC pathophysiology.
Objective: This study aims to evaluate the association between single-nucleotide polymorphisms in MICA rs2596542 and MTHFR rs1801133 and the risk of HCC development.
Methods: A total of 240 participants were enrolled, including 80 individuals with HCC associated with hepatitis C virus (HCV) in Group I, 80 individuals with HCC linked to hepatitis B virus (HBV) in Group II, and 80 age- and gender-matched healthy controls in Group III. Genetic polymorphisms of MICA rs2596542 and MTHFR rs1801133 were identified using real-time polymerase chain reaction analysis. This study is distinct in analyzing these SNPs in a well characterized Egyptian cohort with explicit virus stratified comparisons (HBV vs HCV) and combined genotype–clinical modeling to evaluate both susceptibility and clinical correlates.
Results: A significant increase in the heterozygous CT genotype of MICA rs2596542 and in the T allele frequency was observed in Group II (HCC/HBV) compared to Group III (controls). The MICA CT genotype (rs2596542) was clinically associated with HCC stage and alpha-fetoprotein (AFP) levels. Similarly, the AA and GA genotypes of MTHFR rs1801133 and their mutant A allele frequency were notably elevated in Groups I and II compared to controls. The GA and AA genotypes of MTHFR rs1801133 showed clinical correlations with HCC staging and AFP levels. Multivariate analysis identified hypercalcemia, back pain, metastatic disease, and multiple lesions as independent predictors of survival.
Conclusion: MICA (rs2596542) and MTHFR (rs1801133) polymorphisms were determined to be independent risk factors for HCC disease susceptibility. Notably, they were significantly associated with HCC development. These results indicate that MICA (rs2596542) and MTHFR (rs1801133) could represent valuable markers to recognize individuals who are at a greater probability of developing HCC. The most independent factors affecting survival were hypercalcemia, back pain, metastatic disease, and the occurrence of more than one lesion when multivariate analysis was performed.
Introduction
Hepatocellular carcinoma (HCC) is the most common primary liver malignancy and represents a significant global health burden. Ranking as the sixth most frequently diagnosed cancer worldwide, HCC is also the third leading cause of cancer-related mortality, with a five-year survival rate of less than 20% [1]. The incidence of HCC continues to rise, with projections estimating a 41% increase in global mortality by 2040 [2]. This trend underscores the urgent need for improved early detection methods and targeted therapeutic strategies. In Egypt, HCC has emerged as the second most prevalent cancer among males, accounting for approximately 8.2% of cancer-related deaths [3].
The development of HCC is strongly influenced by various risk factors, including chronic viral infections such as hepatitis B virus (HBV) and hepatitis C virus (HCV), excessive alcohol consumption, diabetes mellitus, and nonalcoholic fatty liver disease (NAFLD) [4]. These factors contribute to chronic liver inflammation, fibrosis, and cirrhosis, creating a microenvironment conducive to tumor initiation and progression. Despite advancements in screening and treatment modalities, the prognosis of HCC remains poor, primarily due to late-stage diagnosis and limited curative options. Early detection significantly improves patient outcomes, as small, localized tumors may be treated with curative interventions such as surgical resection, liver transplantation, or ablative therapies [5].
Recent research has focused on genetic predisposition as a crucial factor in HCC susceptibility. Among the genetic variants associated with HCC risk, the major histocompatibility complex class I chain-related A (MICA) and methylenetetrahydrofolate reductase (MTHFR) genes have garnered considerable attention. The MICA gene encodes a highly polymorphic stress-induced antigen that is expressed on the cell surface, facilitating immune recognition. The protein product of MICA plays a vital role in activating intestinal epithelial gamma delta T cells, which contribute to immune surveillance and tumor suppression [6]. However, genetic variations, such as the MICA rs2596542 single-nucleotide polymorphism (SNP), have been linked to alterations in immune function and tumor development.
The MTHFR gene, located on chromosome 1p36.3, encodes a key enzyme involved in folate metabolism. MTHFR mediates the conversion of 5,10-methylenetetrahydrofolate (5,10-methylene-THF) to 5-methyltetrahydrofolate (5-methyl-THF), a process essential for DNA synthesis and methylation. The MTHFR rs1801133 polymorphism has been extensively studied for its potential role in malignancy, as alterations in folate metabolism can disrupt genomic stability and contribute to tumorigenesis [7]. Several studies have proposed a correlation between this SNP and multiple tumor types, including HCC [8, 9].
Given the rising incidence of HCC and the increasing recognition of genetic influences in cancer susceptibility, further exploration of MICA rs2596542 and MTHFR rs1801133 polymorphisms is warranted. We therefore performed a case–control study in an Egyptian population with explicit stratification by viral etiology (HBV vs HCV) and integrated genotype–clinical analyses to evaluate whether MICA rs2596542 and MTHFR rs1801133 are associated with HCC susceptibility, disease stage, and survival in a population underrepresented in prior meta analyses.
Materials and Methods
This study was conducted on 240 participants in collaboration with the Chemistry Department, Faculty of Science; Clinical Biochemistry Department, Faculty of Medicine; Menoufia; Clinical Oncology and Nuclear Medicine Department, Faculty of Medicine, Menoufia University; and Medical Biochemistry and Molecular Biology Department, Faculty of Medicine, Cairo University. The study protocol was approved by the Ethics Committee of Medical Research at Menoufia University, and informed written consent was obtained from all participants. This study was conducted in accordance with the principles outlined in the Declaration of Helsinki, ensuring ethical standards in medical research involving human subjects.
The subjects were classified into three groups: Group I (n = 80): Patients diagnosed with hepatocellular carcinoma (HCC) associated with hepatitis C virus (HCV). Group II (n = 80): Patients diagnosed with HCC linked to hepatitis B virus (HBV). Group III (n = 80): Healthy age- and gender-matched individuals serving as the control group. All participants underwent comprehensive medical assessments, including detailed history taking, physical examinations, and laboratory investigations. These investigations included liver function tests, HCV antibody screening, HBsAg testing, alpha-fetoprotein (AFP) measurement, and genetic analysis of MICA rs2596542 and MTHFR rs1801133 single-nucleotide polymorphisms (SNPs) from DNA extracted from blood samples using real-time polymerase chain reaction (PCR).
HCC diagnosis was based on elevated AFP levels, imaging findings from triphasic computed tomography (CT) scans, abdominal ultrasonography (US), or magnetic resonance imaging (MRI) conforming to HCC diagnostic criteria, and histopathological confirmation when available. Participants with coexisting HCV and HBV infections, previous liver transplantation, prior HCC treatment, other malignancies, HIV infection, diabetes, renal insufficiency, or thyroid disorders were excluded from this study.
Sample Collection and Laboratory Procedures
Venous blood samples (6 mL) were collected from all participants under strict aseptic conditions. Three mL was placed in an EDTA-containing tube for DNA extraction and stored at -80°C until further processing. Another 3 mL was collected in a plain tube, centrifuged, and stored at -20°C for laboratory investigations.
Liver function tests (ALT and AST) were performed using the Beckman Coulter AU680 analyzer (Beckman Coulter, California, USA) with respective assay kits. HCV antibody screening, HBsAg testing, and AFP quantification were conducted using ElectroChemiLuminescence (ECLA) technology on a cobas immunoassay analyzer.
Allelic discrimination genotyping was employed to analyze SNP variations in the MICA rs2596542 and MTHFR rs1801133 genes. Genotyping was performed in two sequential steps: DNA extraction from whole blood and real-time PCR amplification.
DNA was isolated using the Thermo Scientific Gene JET Genomic DNA Purification Kit (Cat# K0721, manufactured in Lithuania).
Genotyping was conducted using the Applied Biosystems TaqMan Master Mix II (2×), probes, and primers (Foster City, USA, 2010). The manufacturer provided specific probes for SNP analysis:
- M I C A r s 2 5 9 6 5 4 2 [ V I C / F A M ] : TTTTAATGACAACATAGAATTTTAT [C/T] GTGTAGCTGTTCTTTGGGAGACGAT
- M T H F R rs1801 133 [VIC/F A M ] : GAAAAGCTGCGTGATGATGAAATCG [G/A] CTCCCGCAGACACCTTCTCCTTCAA
PCR reactions were prepared with a total volume of 25 µL, comprising 1.25 µL of SNP assay, 12.5 µL of Master Mix, 6.25 µL of nuclease-free water, 5 µL of DNA template for unknown samples, and 5 µL of DNase-free water for negative control reactions. Prior to amplification, all solutions were carefully centrifuged and gently vortexed to ensure homogeneity. The genotyping process was carried out using the Applied Biosystems 7500 real-time PCR system, following a defined thermal cycling protocol. The reaction commenced with an initial denaturation step at 94°C for 4 minutes to separate the DNA strands, followed by 50 amplification cycles. Each cycle consisted of denaturation at 94°C for 30 seconds, annealing at 50°C for 25 seconds to facilitate primer binding, and extension at 72°C for 40 seconds for DNA synthesis. A final extension phase at 72°C was performed for 3 minutes to ensure complete amplification of the target sequences. The Applied Biosystems 7500 PCR system v2.0.1 was employed for DNA amplification, plate documentation, and subsequent analysis of the results.
Amplification reactions were analyzed using Applied BioSystems 7500 PCR system v2.0.1 for result interpretation, plate documentation, and SNP identification (Figures 1 and 2).
Figure 1. Allelic Discrimination Plot. SNP Genotyping Scatter Plot of MICA (rs2596542).
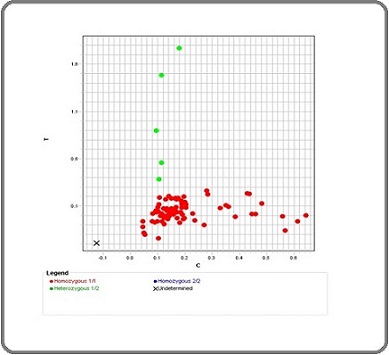
Figure 2. Allelic Discrimination Plot. SNP Genotyping Scatter Plot of MTHFR (rs1801133).
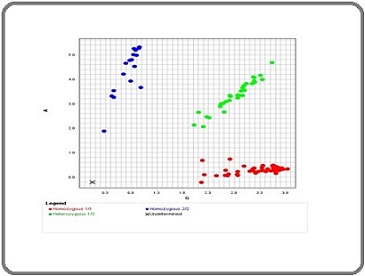
Hardy–Weinberg equilibrium (HWE) was assessed in the control group for each SNP using PLINK/SNPStats. HWE testing was used as a quality control check rather than as a formal test of association. Genotyping quality control measures (per SNP call rates >95%, duplicate concordance ≥99%, and manual review of ambiguous clusters) were applied; SNPs or samples failing QC were re run or excluded. If a SNP was monomorphic in controls (no observed minor allele), HWE cannot be meaningfully assessed.
Statistical analysis
Statistical analysis was conducted using IBM SPSS Statistics software version 20.0 (Armonk, NY: IBM Corp). Qualitative data were presented as frequencies and percentages, while quantitative variables were described using measures such as range (minimum and maximum values), median, interquartile range (IQR), mean, and standard deviation. The Kolmogorov–Smirnov test was employed to assess the normality of data distribution, and statistical significance was determined at a threshold of p < 0.05. The chi-square test was utilized to compare categorical variables among different groups, with corrections applied using Fisher’s Exact test or Monte Carlo correction when more than 20% of expected cell counts were below five. For comparisons involving normally distributed quantitative variables across more than two groups, the F-test (ANOVA) was performed. In cases where quantitative variables exhibited non-normal distribution, the Mann–Whitney test was employed for comparisons between two groups, while the Kruskal–Wallis test was applied for analyzing more than two groups, followed by Dunn’s multiple comparisons test for post hoc pairwise analysis. Additionally, the Hardy–Weinberg equation was used to assess whether the sample population conformed to genetic equilibrium.
We performed virus stratified genotype comparisons (HBV vs controls; HCV vs controls), multivariable logistic regression adjusting for age, sex, cirrhosis, and key laboratory covariates to test independent associations with HCC status, and interaction tests to assess whether genotype effects differed by viral etiology. For survival, we included genotypes and clinical covariates in multivariable Cox models to evaluate independent prognostic effects.
Power calculation was performed to guide sample size and interpret negative findings. With 80 cases per disease group (HCC/HBV and HCC/HCV) and 80 controls, a two sided α = 0.05 provides approximately 80% power to detect an odds ratio (OR) of ~2.0 for a SNP with a minor allele frequency (MAF) of ~0.20 under a log additive model. For SNPs with lower MAF or for smaller effect sizes (OR < 1.7), power is substantially reduced. For survival analyses, assuming an overall event rate of ~50% among 160 cases, the study has roughly 80% power to detect a hazard ratio (HR) of ~1.7–1.9 for a binary predictor; power to detect smaller HRs is limited. These calculations are approximate and were used to interpret both positive and null findings; we report effect estimates and 95% confidence intervals throughout to allow assessment of precision.
Results
This study included a total of 240 participants, comprising 160 cases diagnosed with hepatocellular carcinoma (HCC) and 80 healthy, age- and gender-matched controls. The subjects were categorized into three groups: Group I consisted of 80 individuals with HCC associated with hepatitis C virus (HCV), Group II included 80 individuals with HCC linked to hepatitis B virus (HBV), and Group III served as the control group with 80 healthy participants. We present virus stratified genotype analyses and combined genotype–clinical models: MICA rs2596542 associations were most pronounced in HBV related HCC, whereas MTHFR rs1801133 associations were observed across both viral etiologies; multivariable models adjusting for clinical covariates confirmed the independent association of MTHFR genotypes with case status but not with survival.
The mean age of participants in Groups I, II, and III was 62.81, 62.28, and 63.91 years, respectively. Cirrhosis and clinical symptoms indicative of HCC were observed in 85% of cases within Group I and 30% of cases in Group II (Table 1 and 2).
| Group I HCC/HCV (n = 80) | Group II HCC/HBV (n = 80) | Group III (n = 80) | Test of sig. | P | ||||
| No. | % | No. | % | No. | % | |||
| Gender | ||||||||
| Male | 42 | 52.5 | 46 | 57.5 | 40 | 50 | c 2 = 0.938 | 0.626 |
| Female | 38 | 47.5 | 34 | 42.5 | 40 | 50 | ||
| Age (years) | ||||||||
| Min. – Max. | 50.0 – 77.0 | 50.0 – 76.0 | 47.0 – 75.0 | F= 1.115 | 0.33 | |||
| Mean ± SD. | 62.81 ± 5.79 | 62.28 ± 5.78 | 63.91 ± 9.11 | |||||
| Median (IQR) | 63.50 (58.5 – 66.0) | 63.0 (58.0 – 67.0) | 66.0 (55.0 – 73.0) |
Χ2: Chi square test; F: F for One way ANOVA test; p: p value for comparing between the three studied groups; IQR: Inter quartile range; SD: Standard deviation
| Group I | Group II | χ 2 | p | |||
| HCC/HCV | HCC/HBV | |||||
| (n = 80) | (n = 80) | |||||
| No. | % | No. | % | |||
| History bilhaziasis | 18 | 22.5 | 12 | 15 | 1.477 | 0.224 |
| Smoking | 20 | 25 | 24 | 30 | 0.502 | 0.479 |
| Cirrhosis | 68 | 85 | 24 | 30 | 49.514 * | <0.001 * |
| PVT | 44 | 55 | 40 | 50 | 0.401 | 0.527 |
| Portal hypertension | 34 | 42.5 | 36 | 45 | 0.102 | 0.75 |
| Presenting symptoms | ||||||
| Weight loss | 6 | 7.5 | 6 | 7.5 | 0 | 1 |
| Jaundice | 8 | 10 | 8 | 10 | 0 | 1 |
| Hypercalcemia | 6 | 7.5 | 0 | 0 | 6.234 * | FEp=0.028 * |
| Fracture | 2 | 2.5 | 2 | 2.5 | 0 | FEp=1.000 |
| Follow up | 10 | 12.5 | 28 | 35 | 11.182 * | 0.001 * |
| Dyspnea | 2 | 2.5 | 4 | 5 | 0.693 | FEp=0.681 |
| Cord compression | 2 | 2.5 | 2 | 2.5 | 0 | FEp=1.000 |
| Back pain | 8 | 10 | 8 | 10 | 0 | 1 |
| Abdominal pain/ Dyspepsia | 36 | 45 | 22 | 27.5 | 5.301 * | 0.021 * |
| Antiviral therapy | 40 | 50 | 34 | 42.5 | 0.905 | 0.341 |
| Local ablative treatment | 32 | 40 | 30 | 37.5 | 0.105 | 0.746 |
| Systemic therapy | 40 | 50 | 38 | 47.5 | 0.1 | 0.752 |
| Tumor size | ||||||
| <5 | 38 | 47.5 | 36 | 45 | 0.101 | 0.751 |
| >5 | 42 | 52.5 | 44 | 55 | ||
| Tumor site | ||||||
| Right lobe | 62 | 77.5 | 62 | 77.5 | 0.468 | 0.792 |
| Left lobe | 12 | 15 | 10 | 12.5 | ||
| Right & left lobe | 6 | 7.5 | 8 | 10 | ||
| Tumor number | ||||||
| 1 | 26 | 32.5 | 32 | 40 | 0.974 | 0.324 |
| >1 | 54 | 67.5 | 48 | 60 | ||
| Metastatic site | ||||||
| No | 38 | 47.5 | 42 | 52.5 | 0.735 | 0.972 |
| Bone | 26 | 32.5 | 22 | 27.5 | ||
| Lung | 8 | 10 | 8 | 10 | ||
| LN | 6 | 7.5 | 6 | 7.5 | ||
| Multiple sites | 2 | 2.5 | 2 | 2.5 | ||
| Stage | ||||||
| Early | 24 | 30 | 28 | 35 | 0.543 | 0.762 |
| Metastatic | 36 | 45 | 32 | 40 | ||
| Advanced | 20 | 25 | 20 | 25 |
χ2: Chi square test; FE: Fisher Exact; PVT: Portal vein thrombosis. p: p value for comparing between the two studied groups *: Statistically significant at p ≤ 0.05
A comparative analysis between the patient groups and healthy controls revealed significant reductions in platelet count and albumin levels in Groups I and II, whereas total bilirubin concentration was significantly higher in these groups compared to the control group. Furthermore, levels of alanine aminotransferase (ALT), aspartate aminotransferase (AST), and alpha-fetoprotein (AFP) were markedly elevated in Groups I and II, while prothrombin concentration was significantly lower compared to Group III (Table 3).
| Group I | Group II | Group III | H | p | Sig. bet. grps. | |||
| (n = 80) | (n = 80) | (n = 80) | I vs. II | I vs. III | II vs. III | |||
| Serum creatinine(mg/dl) | ||||||||
| Min. – Max. | 0.60 – 1.70 | 0.70 – 1.60 | 0.70 – 1.30 | 6.304 | 0.043 * | 0.107 | 0.389 | 0.013 * |
| Mean ± SD. | 1.05 ± 0.22 | 1.11 ± 0.24 | 1.01 ± 0.17 | |||||
| Median (IQR) | 1.0 (0.90 – 1.20) | 1.10 (0.90 – 1.30) | 1.0 (0.90 – 1.20) | |||||
| Platelet count (×10 3 /ml) | ||||||||
| Min. – Max. | 59.0 – 355.0 | 40.0 – 345.0 | 178.0 – 400.0 | 97.836 * | <0.001 * | 0.148 | <0.001 * | <0.001 * |
| Mean ± SD. | 168.0 ± 69.21 | 149.94 ± 70.50 | 263.01 ± 50.0 | |||||
| Median (IQR) | 165 (110.5–210.0) | 133.0 (94–201) | 256.0 (230–300) | |||||
| Total bilirubin(mg/dl) | ||||||||
| Min. – Max. | 0.70 – 9.0 | 0.70 – 7.50 | 0.20 – 1.10 | 135.683 * | <0.001 * | 0.777 | <0.001 * | <0.001 * |
| Mean ± SD. | 2.75 ± 2.20 | 2.47 ± 1.68 | 0.61 ± 0.27 | |||||
| Median (IQR) | 2.0 (1.4–3.0) | 1.90 (1.40–3.0) | 0.58 (0.40 – 0.90) | |||||
| Serum albumin(g/dl) | ||||||||
| Min. – Max. | 1.70 – 4.0 | 2.0 – 3.90 | 3.40 – 5.0 | 148.817 * | <0.001 * | 0.759 | <0.001 * | <0.001 * |
| Mean ± SD. | 2.94 ± 0.48 | 2.92 ± 0.43 | 4.32 ± 0.54 | |||||
| Median (IQR) | 3.0 (2.7 – 3.15) | 2.90 (2.70 – 3.15) | 4.25 (4.0 – 4.90) | |||||
| ALT (U/L) | ||||||||
| Min. – Max. | 17.0 – 142.0 | 23.0 – 140.0 | 11.0 – 43.0 | 101.148 * | <0.001 * | 0.213 | <0.001 * | <0.001 * |
| Mean ± SD. | 68.08 ± 35.94 | 72.38 ± 30.92 | 26.43 ± 8.76 | |||||
| Median (IQR) | 65.50 (34.0 – 89.0) | 76.0 (45.0 – 89.5) | 22.50 (20.5– 34.0) | |||||
| AST (U/L) | ||||||||
| Min. – Max. | 23.0 – 165.0 | 23.0 – 150.0 | 9.0 – 45.0 | 118.479 * | <0.001 * | 0.112 | <0.001 * | <0.001 * |
| Mean ± SD. | 75.75 ± 38.27 | 62.83 ± 30.78 | 27.05 ± 8.59 | |||||
| Median (IQR) | 76.0 (42.0 – 98.0) | 57.0 (39.0 – 77.0) | 24.0 (20.5 – 34.0) | |||||
| PC (μg/mL) | ||||||||
| Min. – Max. | 55.0 – 95.0 | 55.0 – 96.0 | 83.0 – 100.0 | 112.110 * | <0.001 * | 0.613 | <0.001 * | <0.001 * |
| Mean ± SD. | 79.95 ± 10.55 | 79.65 ± 10.30 | 94.55 ± 5.06 | |||||
| Median (IQR) | 83.30 (73.1 – 88.0) | 80.6 (75.0 – 86.75) | 97.0 (90.0 – 98.5) | |||||
| AFP (ng/ml) | ||||||||
| Min. – Max. | 28 – 80343 | 20 – 70986 | 1 – 7 | 159.667 | <0.001 * | 0.807 | <0.001 * | <0.001 * |
| Mean ± SD. | 3842.31±12800.62 | 6371.18±15118.18 | 3.85 ± 1.46 | |||||
| Median (IQR) | 571.5 (159–1450) | 565.5 (164.5–2301.5) | 4 (3.0 – 5.0) |
H: H for Kruskal Wallis test, Pairwise comparison bet. each 2 groups was done using Post Hoc Test (Dunn's for multiple comparisons test); p: p value for comparing between the three studied groups; *: Statistically significant at p ≤ 0.05; IQR: Inter quartile range; SD: Standard deviation; PC: prothrombinconcentration; AFP: Alpha fetoprotein; ALT: Alanine transaminase; AST: Aspartate transaminase
Tumor-related parameters, including size, site, number of lesions, presence of metastases, and staging, did not exhibit statistically significant differences between Group I and Group II. Genetic analysis revealed notable findings regarding single nucleotide polymorphisms (SNPs). A significant increase in the CT genotype of MICA rs2596542 and its mutant T allele frequency was observed in Group II (HCC/HBV) compared to Group III (controls). No TT genotypes were observed in any group for MICA rs2596542; the control group was monomorphic (all CC), which limits allele based comparisons. The MICA CT genotype (rs2596542) was clinically associated with HCC stage and AFP levels. Similarly, the AA and GA genotypes of MTHFR rs1801133 and their mutant A allele frequency were notably elevated in Groups I and II compared to controls. The GA and AA genotypes of MTHFR rs1801133 showed clinical correlations with HCC staging and AFP levels (Table 2, Table 4-6).
| Group I | Group II | Group III | χ 2 | MCp | Sig. bet. grps. | ||||||
| HCC/HCV | HCC/HBV | Control | |||||||||
| No. | % | No. | % | No. | % | I vs. II | I vs. III | II vs. III | |||
| MICA rs2596542 | (n = 80) | (n = 80) | (n = 80) | ||||||||
| CC | 76 | 95 | 74 | 92.5 | 80 | 100 | 6.534 * | 0.038 * | 0.514 | 0.12 | 0.028 * |
| CT | 4 | 5 | 6 | 7.5 | 0 | 0 | |||||
| TT | 0 | 0 | 0 | 0 | 0 | 0 | |||||
| HW p0 | 0.819 | 0.727 | – | ||||||||
| Allele frequency | (n = 160) | (n = 160) | (n = 160) | ||||||||
| C | 156 | 97.5 | 154 | 96.3 | 160 | 100 | 6.412 * | 0.039 * | 0.52 | 0.123 | 0.030 * |
| T | 4 | 2.5 | 6 | 3.8 | 0 | 0 | |||||
| MTHFR rs1801133 | (n = 80) | (n = 80) | (n = 80) | ||||||||
| GG | 40 | 50 | 37 | 46.3 | 55 | 68.8 | 10.446 * | 0.034 * | 0.866 | 0.037 * | 0.011 * |
| GA | 28 | 35 | 29 | 36.3 | 20 | 25 | |||||
| AA | 12 | 15 | 14 | 17.5 | 5 | 6.3 | |||||
| HW p0 | 0.07 | 0.061 | 0.076 | ||||||||
| Allele frequency | (n = 160) | (n = 160) | (n = 160) | ||||||||
| G | 108 | 67.5 | 103 | 64.4 | 130 | 81.3 | 12.537 * | 0.002 * | 0.555 | 0.005 * | 0.012 * |
| A | 52 | 32.5 | 57 | 35.6 | 30 | 18.8 |
χ2: Chi square test; MC: Monte Carlo; p: p value for comparing between the studied groups; p: p value for comparing between the studied groups HWp0: p value for Chi square for goodness of fit for Hardy-Weinberg equilibrium
| rs2596542 | Test of | P | |||||
| CC | CT | sig. | |||||
| (n = 74) | (n = 6) | ||||||
| No. | % | No. | % | ||||
| Tumor size | |||||||
| <5 | 33 | 44.6 | 3 | 50 | Χ 2 = 0.066 | FE p =1 | |
| >5 | 41 | 55.4 | 3 | 50 | |||
| Stage | |||||||
| Early | 28 | 37.8 | 0 | 0 | Χ 2 =9.062 * | MC p =0.005 * | |
| Metastatic | 31 | 41.9 | 1 | 16.7 | |||
| Advanced | 15 | 20.3 | 5 | 83.3 | |||
| Serum creatinine | |||||||
| Min. – Max. | 0.70 – 1.60 | 0.0 – 1.30 | U= 157.50 | 0.235 | |||
| Mean ± SD. | 1.12 ± 0.24 | 1.0 ± 0.18 | |||||
| Median | 1.1 | 0.95 | |||||
| Platelet count | |||||||
| Min. – Max. | 40.0 – 345.0 | 93.0 – 230.0 | U= 204.00 | 0.742 | |||
| Mean ± SD. | 151.1 ± 71.88 | 135.67 ± 53.18 | |||||
| Median | 133 | 116 | |||||
| Total bilirubin | |||||||
| Min. – Max. | 0.70 – 7.50 | 0.90 – 7.50 | U= 208.500 | 0.805 | |||
| Mean ± SD. | 2.46 ± 1.63 | 2.61 ± 2.43 | |||||
| Median | 1.95 | 1.8 | |||||
| Serum albumin | |||||||
| Min. – Max. | 2.0 – 3.90 | 2.70 – 3.50 | U= 193.500 | 0.601 | |||
| Mean ± SD. | 2.91 ± 0.43 | 3.03 ± 0.31 | |||||
| Median | 2.9 | 2.95 | |||||
| AFP | |||||||
| Min. – Max. | 20.0 – 70986.0 | 46.0 – 425.0 | U= 99.50 * | 0.025 * | |||
| Mean ± SD. | 6872.32 ± 15618.87 | 190.33 ± 143.10 | |||||
| Median | 81.5 | 175.5 |
Χ2: Chi square test MC: Monte Carlo; FE: Fisher Exact U: Mann Whitney test ; AFP: Alpha fetoprotein
| rs1801133 | |||||||||||||
| group I (HCC/HCV) | groupII (HCC/HBV) (n = 80) | ||||||||||||
| GG | GA | AA | GG | GA | AA | ||||||||
| (n = 40) | (n = 28) | (n = 12) | (n = 37) | (n = 29) | (n = 14) | ||||||||
| No. | % | No. | % | No. | % | No. | % | No. | % | No. | % | ||
| Tumor size | |||||||||||||
| <5 | 20 | 50 | 11 39 | .3 | 7 | 58.3 | 11 | 29.7 | 18 | 62.1 | 7 | 50 | |
| >5 | 20 | 50 | 17 60 | .7 | 5 | 41.7 | 26 | 70.3 | 11 | 37.9 | 7 | 50 | |
| Χ 2 (p) | 1.423 (0.491) | 7.041 * (0.030 * ) | |||||||||||
| Stage | |||||||||||||
| Early | 17 | 42.5 | 5 17 | .9 | 2 | 16.7 | 9 | 24.3 | 14 | 48.3 | 5 | 35.7 | |
| Metastatic | 12 | 30 | 14 | 50 | 10 | 83.3 | 12 | 32.4 | 12 | 41.4 | 8 | 57.1 | |
| Advanced | 11 | 27.5 | 9 32 | .1 | 0 | 0 | 16 | 43.2 | 3 | 10.3 | 1 | 7.1 | |
| Χ 2 (MC P ) | 13.594 (0.007 * ) | 12.751 * (0.011 * ) | |||||||||||
| Serum creatinine | |||||||||||||
| Min. – Max. | 0.60 – 1.70 | 0.80 – 1.50 | 0.80 – 1.70 | 0.70 – 1.60 | 0.80 – 1.50 | 0.80 – 1.50 | |||||||
| Mean ± SD. | 1.03 ± 0.23 | 1.10 ± 0.21 | 1.01 ± 0.24 | 1.12 ± 0.26 | 1.15 ± 0.21 | 1.01 ± 0.22 | |||||||
| Median | 1 | 1.1 | 0.95 | 1.1 | 1.2 | 0.95 | |||||||
| H (p) | 2.608 (0.272) | 3.776 (0.151) | |||||||||||
| Platelet count | |||||||||||||
| Min. – Max. | 59.0 – 355.0 | 78.0 – 345.0 | 94.0 – 281.0 | 40.0 – 230.0 | 60.0 – 230.0 | 61.0 – 345.0 | |||||||
| Mean ± SD. | 162.8 ± 73.52 | 169.1 ± 66.88 | 182.7 ± 62.63 | 144.9 ± 62.01 | 145.2 ± 56.43 | 172.9 ± 109.6 | |||||||
| Median | 178 | 160.5 | 158.5 | 133 | 145 | 128 | |||||||
| H(p) | 1.152 (0.562) | 0.059 (0.971) | |||||||||||
| Total bilirubin | |||||||||||||
| Min. – Max. | 0.80 – 9.0 | 0.70 – 8.0 | 1.0 – 3.0 | 0.80 – 7.50 | 0.70 – 7.0 | 0.70 – 5.0 | |||||||
| Mean ± SD. | 3.32 ± 2.72 | 2.35 ± 1.51 | 1.81 ± 0.67 | 2.81 ± 1.86 | 2.11 ± 1.46 | 2.33 ± 1.54 | |||||||
| Median | 2 | 2.1 | 1.75 | 2 | 1.7 | 1.85 | |||||||
| H (p) | 1.616 (0.446) | 2.814 (0.245) | |||||||||||
| Serum albumin | |||||||||||||
| Min. – Max. | 1.70 – 3.80 | 2.0 – 4.0 | 1.80 – 3.60 | 2.0 – 3.90 | 2.0 – 3.80 | 2.30 – 3.80 | |||||||
| Mean ± SD. | 2.88 ± 0.45 | 3.04 ± 0.48 | 2.88 ± 0.58 | 2.93 ± 0.45 | 2.92 ± 0.42 | 2.89 ± 0.41 | |||||||
| Median | 2.9 | 3 | 3 | 3 | 2.9 | 2.9 | |||||||
| H (p) | 2.185 (0.335) | 0.196 (0.907) | |||||||||||
| AFP | |||||||||||||
| Min. – Max. | 28.0 – 19109.0 | 38.0 – 80343.0 | 92.0 – 7323.0 | 20.0 – 70986.0 | 27.0 – 56432.0 | 88.0 – 55467.0 | |||||||
| Mean ± SD. | 1576.2 ± 3496.7 | 7753.5± 20835.7 | 2270.1 ± 2745.6 | 9660.62±17807.98 | 2864.14± 10447.93 | 4942.21±14669.80 | |||||||
| Median | 333.5 | 1096 | 555 | 1330 | 233 | 555 | |||||||
| H (p) | 6.413 (0.040 * ) | 4.671 (0.097) |
χ2: Chi square test; MC: Monte Carlo; H: H for Kruskal Wallis test; p: p value for comparison between the studied categories; *: Statistically significant at p ≤ 0.05; SD: Standard deviation; AFP: Alpha fetoprotein
Hardy–Weinberg equilibrium (HWE) was evaluated for each polymorphism in all study groups. For MICA rs2596542, genotype distributions in the HCC/HCV group (HW p = 0.819) and the HCC/HBV group (HW p = 0.727) were consistent with HWE, indicating no evidence of genotyping error or population stratification within these groups. In the control group, all individuals carried the CC genotype, resulting in a monomorphic distribution; therefore, deviation from HWE could not be meaningfully assessed. For rs1801133, genotype frequencies were in Hardy–Weinberg equilibrium in the HCC/HCV group (HW p = 0.070), the HCC/HBV group (HW p = 0.061), and the control group (HW p = 0.076), suggesting reliable genotyping and absence of significant population stratification across all studied groups (Table 4).
Univariate analysis identified several factors influencing survival outcomes, including portal vein thrombosis (PVT), weight loss, hypercalcemia, dyspnea, and back pain, all of which had a significant impact on survival. In addition, advanced tumor stage, larger tumor size, presence of multiple lesions, and type of treatment administered were also significant prognostic determinants (Table 7).
| Univariate | #Multivariate | |||
| P | HR (LL – UL 95%C.I) | p | HR (LL – UL 95%C.I) | |
| Sex (female) | 0.306 | 1.258 (0.811 – 1.952) | ||
| Age (years) | 0.144 | 0.971 (0.932 – 1.010) | ||
| HCVAb | 0.165 | 1.367 (0.879 – 2.124) | ||
| HBsAg | 0.165 | 0.732 (0.471 – 1.137) | ||
| History bilhaziasis | 0.695 | 0.891 (0.500 – 1.586) | ||
| Smoking | 0.083 | 0.629 (0.372 – 1.063) | ||
| Cirrhosis | 0.439 | 1.196 (0.760 – 1.881) | ||
| PVT | <0.001 * | 2.678 (1.657 – 4.330) | 0.447 | 1.323 (0.643–2.722) |
| Portal hypertension | 0.786 | 1.063 (0.684 – 1.651) | ||
| Presenting symptoms | ||||
| Weight loss | 0.027* | 2.123 (1.089 – 4.138) | 0.489 | 1.362 (0.567–3.271) |
| Jaundice | 0.354 | 1.338 (0.723 – 2.476) | ||
| Hypercalcemia | <0.001 * | 11.492 (4.578 – 28.848) | 0.001 * | 7.757 (2.394–25.131) |
| Fracture | 0.055 | 2.703 (0.978 – 7.473) | ||
| Dyspnea | 0.004 * | 3.440 (1.486 – 7.965) | 0.486 | 1.487 (0.487–4.541) |
| Cord compression | 0.197 | 1.944 (0.707 – 5.341) | ||
| Back pain | <0.001 * | 2.901 (1.620 – 5.195) | 0.009* | 3.394 (1.353–8.513) |
| Abdominal pain/ Dyspepsia | 0.296 | 0.775 (0.480 – 1.250) | ||
| Antiviral therapy | 0.001 * | 0.441 (0.275 – 0.706) | 0.979 | 1.009 (0.500–2.036) |
| Local ablative treatment | <0.001 * | 0.050 (0.018 – 0.136) | 0.147 | 0.389 (0.109–1.392) |
| Systemic therapy | <0.001 * | 3.197 (1.995 – 5.123) | 0.488 | 1.243 (0.672–2.297) |
| Tumor size (>5) | <0.001 * | 6.161 (3.439 – 11.040) | 0.314 | 0.679 (0.319–1.444) |
| Tumor number (>1) | <0.001 * | 17.907 (6.525 – 49.146) | 0.001* | 14.775 (3.136–69.622) |
| Metastatic | <0.001 * | 23.663 (11.124–50.336) | <0.001 * | 22.908 (5.442–96.427) |
| Stage (Advanced) | 0.002 * | 0.373 (0.202 – 0.691) | 0.398 | 0.596 (0.179–1.979) |
| Serum creatinine | 0.502 | 1.378 (0.540 – 3.519) | ||
| Platelet count | 0.235 | 0.998 (0.995 – 1.0) | ||
| Total bilirubin | <0.001 * | 1.186 (1.085 – 1.295) | 0.087 | 1.132 (0.982–1.305) |
| Serum albumin | 0.001 * | 0.457 (0.285 – 0.731) | 0.244 | 0.681 (0.357–1.300) |
| ALT | <0.001 * | 1.015 (1.018 – 1.022) | 0.548 | 0.997 (0.987–1.007) |
| AST | 0.003 * | 1.009 (1.003 – 1.015) | 0.381 | 1.004 (0.995–1.012) |
| PT | 0.96 | 1.001 (0.979 – 1.022) | ||
| AFP | <0.001 * | 1.0 (1.0 – 1.0) | 0.923 | 1.0 (1.0 – 1.0) |
| rs2596542 (CT) | 0.621 | 0.776 (0.284 – 2.121) | ||
| rs1801133 (GA + AA) | 0.472 | 1.175 (0.757 – 1.823) |
The mean overall survival was 14.6 months for Group I and 16.3 months for Group II, with no statistically significant difference between the two groups (p = 0.258) (Table 8, Figure 3).
| Mean | % End of study | Log rank | ||
| χ 2 | p | |||
| Group I | 14.661 | 44.9 | 1.278 | 0.258 |
| Group II | 16.265 | 54.9 |
Figure 3. Kaplan-Meier Survival Curve for Overall Survival.
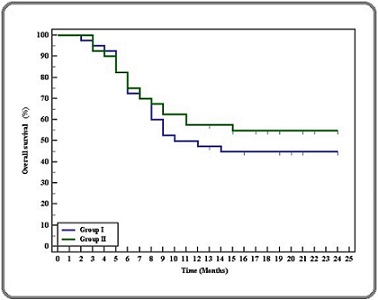
Neither rs2596542 nor rs1801133 was significantly associated with overall survival in multivariable models. Given the sample size and the number of events, the study had limited power to detect small to moderate genotype effects on. Therefore, absence of a statistically significant association should be interpreted cautiously; the 95% confidence intervals reported for hazard ratios indicate the range of effect sizes compatible with our data. Multivariate analysis further refined these observations, identifying hypercalcemia, back pain, metastatic disease, and the presence of multiple lesions as the most independent survival predictors (p-values = 0.001, 0.009, 0.001, and <0.001, respectively). Notably, the polymorphisms rs2596542 and rs1801133 did not exhibit a statistically significant impact on survival outcomes or treatment selection (p = 0.621 and 0.472, respectively) (Table 7).
Discussion
This study investigated the correlation between MICA rs2596542 and MTHFR rs1801133 gene single nucleotide polymorphisms (SNPs) with hepatocellular carcinoma (HCC) risk. The study groups were frequency matched for key demographic variables to reduce confounding. The major histocompatibility complex class I related chain A (MICA) gene, located on chromosome 6p21.33, has been widely studied for its tumor specific antigen properties and extensive polymorphisms [10, 11]. MICA encodes a stress induced ligand for the activating receptor NKG2D on NK cells and cytotoxic T lymphocytes; altered expression or proteolytic shedding of MICA can impair immune surveillance and facilitate tumor immune escape. Another key protein implicated in carcinogenesis is methylenetetrahydrofolate reductase (MTHFR), an enzyme critical to folate metabolism, DNA synthesis, methylation, and repair. The relationship between MTHFR polymorphisms and HCC risk has been explored in previous studies, although conflicting results persist [12].
MICA rs2596542 Genotypes
The present study assessed the association between the MICA rs2596542 variant and clinical parameters in patients with HCC/HCV, HCC/HBV, and controls. We observed a significant increase in the CT genotype and the T allele frequency in HBV related HCC (Group II) compared with controls, whereas no significant difference was detected in HCV related HCC (Group I). The CT genotype correlated with higher AFP and more advanced stage but did not independently predict overall survival after multivariable adjustment.
Etiology specific interpretation and mechanisms HBV and HCV differ in their biology and immune interactions, which may explain the etiology specific association we observed. HBV is a DNA virus capable of genomic integration and expression of viral proteins (e.g., HBx) that modulate host transcription and immune signaling; these processes can alter MICA expression or increase proteolytic shedding of MICA, thereby reducing cell surface ligand availability and impairing NKG2D mediated cytotoxicity. By contrast, HCV establishes chronic inflammation and immune exhaustion through persistent antigenic stimulation and interferon pathway modulation; in this context, chronic immune dysregulation may overshadow modest germline effects on NKG2D ligands. Therefore, rs2596542 may exert stronger functional consequences in HBV infected livers than in HCV infected livers, a hypothesis that requires direct testing.
Comparison with prior studies Several studies and meta analyses have assessed rs2596542, frequently yielding inconsistent conclusions due to variations in ethnicity, sample size, and underlying HCC etiology [8, 13, 14]. Meta analyses suggest that the T allele of rs2596542 may be a risk factor for HCC development, whereas the C allele appears protective [13, 14]. Our virus stratified results indicate that pooling HBV and HCV cases in meta analyses could mask etiology specific effects, which helps reconcile some prior discrepancies.
Recommended mechanistic follow up To resolve conflicting epidemiologic signals, future studies should measure tumor cell surface MICA, circulating soluble MICA (sMICA), and NKG2D expression on immune cells, stratified by viral etiology and rs2596542 genotype. Functional assays assessing transcriptional activity, protein stability, and proteolytic shedding in hepatocytes or hepatoma cell lines expressing HBV or HCV proteins would clarify whether rs2596542 modifies MICA biology differently in HBV versus HCV infection.
MTHFR rs1801133 Genotypes
Our study examined the association between MTHFR rs1801133 polymorphisms and HCC risk. Results demonstrated a statistically significant elevation in the AA and GA genotypes and the A allele frequency in Groups I and II compared to Group III. GA and AA genotypes correlated with more advanced stage and higher AFP.
Biological plausibility and literature context The C677T (Ala222Val) substitution reduces MTHFR enzymatic activity and can perturb folate metabolism, DNA synthesis, and methylation mechanisms that plausibly contribute to genomic instability and carcinogenesis. Associations between MTHFR variants and cancer risk vary by tumor type, population folate status, alcohol exposure, and other environmental modifiers; this complexity likely explains heterogeneous findings across studies [8, 15-18].
Implications Our multivariable analyses indicate that the MTHFR association with HCC case status persists after adjustment for clinical covariates, supporting a potential role in susceptibility within this Egyptian cohort. Future work should evaluate folate biomarkers and alcohol exposure to assess gene–environment interactions that may modify risk.
Treatment Options and Prognostic Factors
In Western populations, alcoholic fatty liver disease and nonalcoholic fatty liver disease have become important HCC etiologies [19, 20]. Systemic treatment options were comparable between Groups I and II, with sorafenib and best supportive care being the most frequently used approaches. Univariate analysis identified several clinical factors associated with poorer survival (portal vein thrombosis, weight loss, hypercalcemia, dyspnea, and back pain), and multivariable Cox regression identified hypercalcemia, metastatic disease, back pain, and multiple lesions as independent predictors.
Immunotherapy limitation and generalizability A key limitation is that none of our patients received contemporary immunotherapy combinations (for example anti PD 1/PD L1 agents with anti VEGF). Immune checkpoint inhibitors substantially remodel the tumor immune microenvironment and can interact with pathways involving NKG2D and its ligands; therefore, prognostic models derived from pre immunotherapy cohorts may not generalize to current treatment settings. Germline variants that affect immune recognition, such as MICA polymorphisms, could plausibly influence response to checkpoint blockade; retrospective or prospective evaluation of these SNPs in immunotherapy treated cohorts is needed to determine predictive value.
Genotyping quality, HWE, and limitations
We assessed Hardy–Weinberg equilibrium as a quality control measure. Case groups were in HWE for the analyzed SNPs; the control group was monomorphic for MICA rs2596542 (all CC), which prevents meaningful HWE testing for that SNP in controls and limits allele based inference. Genotyping QC metrics (per SNP call rates >95% and duplicate concordance ≥99%) support the validity of the observed genotype distributions and argue against systematic assay failure. Nevertheless, the monomorphic control sample and the moderate sample size (80 per group) reduce power for detecting small effects and constrain subgroup analyses. We acknowledge that the control group was monomorphic for MICA rs2596542 in our sample, which limits allele based inference for this SNP; however, genotyping QC metrics and duplicate concordance argue against assay failure, and replication in larger, population representative control cohorts is warranted. Although our cohort is moderate in size and well characterized, the sample size (80 per group) limits power for genetic association analyses, particularly for subgroup analyses (e.g., genotype correlations with AFP or stage) and for detecting modest effects on survival. We therefore emphasize effect estimates and 95% confidence intervals rather than relying solely on p values. Null results for survival should be interpreted as inconclusive for small effect sizes rather than definitive evidence of no effect. Future studies with larger sample sizes, pooled analyses, or meta analyses are warranted to detect smaller genotype effects and to validate our findings.
Recommendations for future research Replication in larger, multi center cohorts with population representative controls is essential. Functional studies should measure cell surface and soluble MICA, NKG2D expression, and folate biomarkers, and should test interactions with viral proteins (HBV and HCV). Evaluating these SNPs in immunotherapy treated cohorts and integrating germline genotyping with tumor immune profiling will clarify predictive and prognostic relevance. In conclusion, in this Egyptian cohort, MICA rs2596542 was associated with HBV related HCC and with markers of disease progression, while MTHFR rs1801133 was associated with HCC susceptibility across viral etiologies and correlated with stage and AFP. These associations are hypothesis generating and require replication and mechanistic validation before clinical application.
Declarations
Funding:
No external funding was received for this study.
Conflicts of interest
The authors declare that they have no competing interests.
Ethics approval
The study protocol was approved by the Ethical Scientific Committee of Menoufia University (reference: 11/2022BIO1 1) and was conducted in accordance with the Declaration of Helsinki.
Consent to participate
Written informed consent was obtained from all participants prior to enrollment.
Consent for publication
All participants provided written consent for anonymized, aggregate data from this study to be published. No individual participant is identifiable from the data or figures presented.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. Additional de identified data are available from the corresponding author on reasonable request, subject to institutional and ethical restrictions to protect participant confidentiality.
Code availability
Not applicable. No custom code was used for the statistical analyses reported in this study.
Author contributions
1. S.G.: Conceptualization; study design; supervision; clinical studies; manuscript drafting; manuscript review and editing. 2. A.A.: Laboratory methodology and genotyping; data acquisition; formal analysis; manuscript drafting and editing. 3. A.E.: Data curation; statistical analysis; preparation of tables and figures; manuscript editing. 4. A.S.: Clinical recruitment and characterization; collection of clinical and oncologic data; interpretation of clinical findings. 5. R.H.: Methodological oversight; critical revision of the manuscript for important intellectual content; final approval of the version to be published.
All authors read and approved the final manuscript.
Use of large language models
A final English language proofreading has been conducted using CoPilot® artificial intelligence model (AI). No generative AI was used in this manuscript.
Acknowledgements
The authors thank the staff of the Clinical Biochemistry and Clinical Oncology departments at Menoufia University and the Medical Biochemistry and Molecular Biology Department at Cairo University for their assistance with sample collection and laboratory work. We also thank the study participants for their contribution.
References
- Conceptual Model for the Hepatocellular Carcinoma Screening Continuum: Current Status and Research Agenda Singal AG , Lok AS , Feng Z, Kanwal F, Parikh ND . Clinical Gastroenterology and Hepatology: The Official Clinical Practice Journal of the American Gastroenterological Association.2022;20(1). CrossRef
- The liver cancer immune microenvironment: Therapeutic implications for hepatocellular carcinoma Donne R, Lujambio A. Hepatology.2023;77(5). CrossRef
- Associations between twelve common gene polymorphisms and susceptibility to hepatocellular carcinoma: evidence from a meta-analysis Quan Y, Yang J, Qin T, Hu Y. World Journal of Surgical Oncology.2019;17(1). CrossRef
- From diagnosis to treatment of hepatocellular carcinoma: An epidemic problem for both developed and developing world Dimitroulis D, Damaskos C, Valsami S, Davakis S, Garmpis N, Spartalis E, Athanasiou A, et al . World Journal of Gastroenterology.2017;23(29). CrossRef
- Should we undertake surveillance for HCC in patients with MAFLD? Norero B, Dufour . Therapeutic Advances in Endocrinology and Metabolism.2023;14. CrossRef
- Major histocompatibility complex (MHC) associations with diseases in ethnic groups of the Arabian Peninsula Al Naqbi H, Mawart A, Alshamsi J, Al Safar H, Tay GK . Immunogenetics.2021;73(2). CrossRef
- DNA methylation: an epigenetic mark of cellular memory Kim M, Costello J. Experimental & Molecular Medicine.2017;49(4). CrossRef
- Methylenetetrahydrofolate reductase C677T (Ala>Val, rs1801133 C>T) polymorphism decreases the susceptibility of hepatocellular carcinoma: a meta-analysis involving 12,628 subjects Zhang S, Jiang J, Tang W, Liu L. Bioscience Reports.2020;40(2). CrossRef
- Epidemiology and Management of Hepatocellular Carcinoma Kulik L, El-Serag HB . Gastroenterology.2019;156(2). CrossRef
- MICA polymorphisms and cancer risk: a meta-analysis Ji M, Wang J, Yuan L, Zhang Y, Zhang J, Dong W, Peng X. International Journal of Clinical and Experimental Medicine.2015;8(1).
- Role of MICA rs2596542 and IFN-gamma rs2069727 Polymorphism in HCC Development in Egyptian Patients with HCV-related HCC Morsy AT . Egyptian Journal of Medical Microbiology.2023. CrossRef
- Methylene tetrahydrofolate reductase (MTHFR) and vascular endothelial growth factor (VEGF) polymorphisms in Brazilian patients with Hepatitis C virus (HCV)-related hepatocellular carcinoma (HCC) Carvalho SCR , Vasconcelos LRS , Fonseca L, Carmo RF , Tomitão MT , Aroucha DCBL , Pereira LMMB , et al . Clinics.2021;76. CrossRef
- Relationship between the rs2596542 polymorphism in the MICA gene promoter and HBV/HCV infection-induced hepatocellular carcinoma: a meta-analysis Luo X, Wang Y, Shen A, Deng H, Ye M. BMC medical genetics.2019;20(1). CrossRef
- Association between MICA rs2596542 Polymorphism with the Risk of Hepatocellular Carcinoma in Chronic Hepatitis C Patients Marangon CG , Bitencorte JT , Michita RT , Lunge VR , Dos Santos DC , Álvares-da-Silva MR , Simon D. Pathology oncology research: POR.2020;26(3). CrossRef
- The MTHFR polymorphism affect the susceptibility of HCC and the prognosis of HCC liver transplantation Wang C., Xie H., Lu D., Ling Q., Jin P., Li H., Zhuang R., Xu X., Zheng S.. Clinical & Translational Oncology: Official Publication of the Federation of Spanish Oncology Societies and of the National Cancer Institute of Mexico.2018;20(4). CrossRef
- Correlation Between MTHFR Polymorphisms and Hepatocellular Carcinoma: A Meta-analysis Su H. Nutrition and Cancer.2019;71(7). CrossRef
- Genetic variations in the one-carbon metabolism pathway genes and susceptibility to hepatocellular carcinoma risk: a case-control study Zhang H, Liu C, Han Y, Ma Z, Zhang H, Ma Y, Liu X. Tumour Biology: The Journal of the International Society for Oncodevelopmental Biology and Medicine.2015;36(2). CrossRef
- Association of MTHFR and TYMS gene polymorphisms with the susceptibility to HCC in Egyptian HCV cirrhotic patients Abdel Allah HMM , Zahran WE , El-Masry SA , El-Bendary M, Soliman AF . Clinical and Experimental Medicine.2022;22(2). CrossRef
- Precision pathology analysis of the development and progression of hepatocellular carcinoma: Implication for precision diagnosis of hepatocellular carcinoma Ueno A, Masugi Y, Yamazaki K, Kurebayashi Y, Tsujikawa H, Effendi K, Ojima H, Sakamoto M. Pathology International.2020;70(3). CrossRef
- Diet, lifestyle factors, comorbidities, and hepatocellular carcinoma risk in a middle eastern country: a case-control study Almohaid S, Akhtar S. BMC cancer.2024;24(1). CrossRef
License

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Copyright
© Asian Pacific Journal of Cancer Biology , 2026
Author Details