Hierlungxai: A Hierarchical and Explainable Deep Learning Framework for Ct-Based Lung Cancer Classification
Download
Abstract
Accurate detection of lung cancer (LC) from Computed Tomography (CT) scans plays a crucial role in early diagnosis and effective treatment planning. Despite significant progress in deep learning (DL) based classification, current approaches continue to face challenges such as inconsistent intensity normalization, variable slice resolution, noise artifacts, limited dataset availability, subtle nodule appearance, and complex inter-slice dependencies. To address the identified limitations, this study suggests a new approach HierLungXAI, a comprehensive framework integrating advanced preprocessing, hierarchical feature extraction, and explainable AI techniques. The Advanced Image Standardization and Enhancement Pipeline (AISEP) standardizes intensities, enhances contrast, reduces noise, preserves critical structural details, and augments the dataset to improve model generalization. For feature extraction, the HierEffNet (Hierarchical Efficient-based Network) combines deep hierarchical convolutional modules for local nodule-specific details with a hierarchical attention mechanism to capture global volumetric context across consecutive slices. Extracted features are classified using a Multi-Layer Perceptron (MLP), while Grad-CAM provides visual explanations, highlighting key regions influencing predictions. The proposed framework was evaluated on two benchmark datasets, achieving an accuracy of 99.78% and 99.94%, demonstrating superior performance over existing methods. The proposed integration of AISEP preprocessing, HierEffNet hierarchical feature extraction, MLP classification, and Grad-CAM-based interpretability represents a novel approach that simultaneously enhances sensitivity to small and subtle nodules while effectively modeling inter-slice and global contextual relationships, establishing a robust and transparent framework for clinically reliable LC detection.
1. Introduction
Lung cancer (LC) remains one of the leading causes of cancer-related mortality worldwide, and early detection plays a critical role in improving survival rates and treatment outcomes [1]. Computed Tomography (CT) imaging is widely regarded as the most effective modality for lung cancer screening, as it provides high-resolution cross-sectional visualization of lung structures and enables differentiation between benign and malignant pulmonary nodules. Consequently, accurate and reliable CT-based nodule classification is of paramount clinical importance, directly influencing diagnostic confidence, therapeutic decision-making, and patient prognosis [2].
Despite notable advances in deep learning (DL)-based approaches, automated CT-based LC detection continues to face several persistent challenges [3]. Variations in scanner types, imaging protocols, slice thickness, and acquisition settings introduce significant inconsistencies across datasets, limiting model generalization. In addition, low-dose CT acquisitions, commonly used to reduce radiation exposure, often suffer from increased noise and reduced contrast, making subtle nodules particularly difficult to detect [4]. These factors collectively degrade feature quality and hinder the robustness of automated diagnostic systems.
Another major challenge lies in the high variability of lung nodules, which can differ substantially in size, shape, texture, and intensity [5]. Many existing methods emphasize either localized slice-level features or global volumetric representations in isolation, resulting in incomplete characterization of nodules. In particular, inadequate modeling of inter-slice dependencies leads to fragmented feature representations, reducing sensitivity to irregular or small nodules that span multiple slices [6]. Data-related limitations further complicate this problem. Publicly available CT datasets are often limited in size and exhibit class imbalance between malignant and normal cases [7]. Such constraints increase the risk of overfitting and restrict model robustness across diverse patient populations. Therefore, effective preprocessing, intensity standardization, and data augmentation strategies are essential to enhance image quality, reduce variability,and improve generalization [8].
Beyond accuracy, interpretability is a critical requirement for clinical adoption. Black-box DL models provide limited insight into the decision-making process, which can undermine radiologists’ trust in automated predictions [9]. Without explainable mechanisms to highlight diagnostically relevant regions, models may inadvertently rely on spurious features. Explainable AI techniques are thus vital to ensure that predictions are grounded in meaningful pathological patterns and to support transparent clinical decision-making [10].
These limitations highlight the need for a unified and systematic framework that simultaneously standardizes CT images, enhances subtle nodule detection, models inter-slice and volumetric context, and provides interpretable outputs for clinical use [11-13].
To address these challenges, this study proposes a comprehensive hierarchical and explainable framework for CT-based lung cancer detection. The primary contributions of this work are summarized as follows:
• A robust preprocessing pipeline that standardizes image intensities, enhances contrast, suppresses noise, and augments data to improve consistency and generalization across CT scans.
• A hierarchical feature extraction strategy that jointly captures fine-grained local nodule characteristics and global volumetric context across consecutive slices.
• An effective classification mechanism that reduces overfitting while improving sensitivity to subtle malignant patterns.
• An explainable AI component that visualizes discriminative regions influencing predictions, enhancing transparency and clinical trust.
The remainder of this paper is organized as follows. Section 2 reviews related work in CT-based lung cancer detection. Section 3 describes the proposed methodology, while Section 4 details the datasets, experimental setup, and evaluation metrics. Section 5 presents and discusses the experimental results, including interpretability analysis and comparative evaluations. Finally, Section 6 concludes the paper and outlines directions for future research.
2. Literature Review
Automated LC identification and categorization from CT scans has greatly improved with recent DL advances. Hammad et al. [14] suggested Grad-CAM, a bespoke CNN combined with XAI, to categorize LC into subtypes such as adenocarcinoma, large cell carcinoma, and squamous cell carcinoma. Their model showed strong performance with high precision, recall, and F1-scores and an overall accuracy of 93.06%. Grad-CAM representations also made clinical decision-making interpretable. However, the model faced challenges in handling high variability in nodule appearance and complex inter-slice dependencies, limiting generalization and volumetric contextual understanding. Golkarieh et al. [15] explored the integration of U-Net architectures with different CNN backbones for lung segmentation and classification on a balanced dataset of 832 CT images. UNet with ResNet50 excelled in cancerous segmentation, whereas UNet with VGG16 achieved superior performance for non-cancerous segmentation. Despite high performance, the use of low-resolution images (128×128 pixels) and the lack of explicit modeling of volumetric dependencies could reduce sensitivity to subtle nodules and overall global lung context.
Similarly, Klangbunrueang et al. [16] employed the VGG16 CNN architecture to classify CT scans into Normal, Benign, and Malignant categories. Among 1,097 images from 110 patients, VGG16 outperformed ResNet50, InceptionV3, and MobileNetV2 with a test accuracy of 98.18%, highlighting its potential as a diagnostic support tool. Nonetheless, the relatively small dataset constrained model generalization to unseen or diverse cases. Saha et al. [17] developed VER-Net, a novel transfer learning framework that stacks three pre-trained models to classify four LC types. Using preprocessing, data augmentation, and hyperparameter tuning, VER- Net achieved 91% accuracy, 92% precision, 91% recall, and 91.3% F1-score, outperforming eight other transfer learning models. However, reliance on pre-trained networks may limit domain-specific feature learning and generalization to highly heterogeneous datasets.
Other volumetric-focused approaches, such as, Eldho et al. [18] introduced a 3D DL CNN (3D-DLCNN) approach combined with 3D Mask-R-CNN for pulmonary nodule detection. The method employed Kernel Density Estimation (KDE) and statistical outlier detection for early error removal, achieving 93% accuracy, 92.7% sensitivity, 93.4% specificity, and a 0.87 C-index. While effective, full 3D volumetric processing increases computational complexity, potentially restricting real-time clinical deployment. Hybrid architectures integrating CNNs and Vision Transformers, as in Ozdemir et al. [19] combining InceptionNeXt blocks with grid and block attention to capture both fine-grained local features and global patterns. Evaluated on Chest CT and IQ-OTH/ NCCD datasets, the model achieved 99.54% and 98.41% accuracy, respectively. Despite its superior performance, the model’s computational complexity may limit real-time clinical applicability Shariff et al. [8] developed a CNN framework integrated with Differential Augmentation (DA) to reduce memory overfitting and improve generalization. By applying targeted augmentations and optimizing via Random Search, the model achieved 98.78% accuracy across multiple datasets. While effective, reliance on extensive augmentation increases computational overhead and may not fully capture volumetric contextual information. Gulsoy et al. [20] introduced FocalNeXt, a hybrid ConvNeXt-FocalNet architecture designed to capture both local and global lung features. Evaluated on the IQ-OTH/NCCD dataset, FocalNeXt achieved 99.81% accuracy, 99.78% sensitivity, 99.36% recall, and 99.56% F1-score. Despite its high performance, the hybrid attention-based model demands significant computational resources, potentially limiting deployment in low-resource clinical settings.
DL-LCD, a DL model for LC detection in CT and chest X-ray images, was proposed by Thanammal et al. [21] using preprocessing and lung segmentation using the Honey Badger Algorithm. The approach achieved 98.5% accuracy for normal lungs and 97.44% for abnormal lungs, surpassing existing methods. However, reliance on 2D slice-based analysis limited the model’s ability to capture inter-slice dependencies and volumetric context. Sen et al. [22] introduced a Stacked Neural Network architecture LC detection and classification. Using segmentation followed by feature extraction and classification, the model achieved 96% accuracy with strong precision, recall, and F1-measure. Similar to DL-LCD, SNN’s dependence on 2D slices restricts volumetric contextual learning and inter-slice feature integration.
Overall, while these studies have demonstrated high accuracy in LC detection and classification, common limitations include: small or imbalanced datasets, inadequate modeling of volumetric context and inter-slice dependencies, reliance on 2D analysis or pre-trained networks, and high computational complexity for real-time deployment. Table 1 summarizes recent studies on DL based LC detection, highlighting the dataset used, and model architecture and identified limitations.
| Study | Model / Method | Key Features | Dataset | Key Limitations |
| Hammad et al. [14] | CNN + Grad-CAM | Grad-CAM for interpretability; subtype classification | CT scans | Difficulty handling high nodule variability & inter-slice dependencies |
| Golkarieh et al. [15] | U-Net + CNN backbones | U-Net for lung segmentation; hybrid CNN classifiers | 832 CT images | Low resolution, lacks volumetric modeling |
| Klangbunrueang et al. [16] | VGG16 | VGG16 for 3-class classification | 1,097 CT images | Small dataset limits generalization |
| Saha et al. [17] | VER-Net (transfer learning) | Stacked transfer learning models; hyperparameter tuning | Multiclass CT | Reliance on pre-trained networks may limit domain-specific learning |
| Eldho et al. [18] | 3D-DLCNN + 3D Mask-RCNN | 3D CNN for volumetric features; KDE for error removal | 355 instances / 251k images | High computational complexity |
| Ozdemir et al. [19] | CNN + ViT | Hybrid CNN + Vision Transformer; multi-scale attention | Chest CT + IQ-OTH/NCCD | Computationally heavy; limits real-time deployment |
| Shariff et al.[8] | CNN + Differential Augmentation | Targeted augmentation strategies; Random Search tuning | IQ-OTH/NCCD | Extensive augmentation adds computational overhead |
| Gulsoy et al. [20] | FocalNeXt | ConvNeXt + FocalNet; attention mechanism for local & global features | IQ-OTH/NCCD | High computational requirements |
| Thanammal et al. [21] | DL-LCD | Honey Badger segmentation; CLAHE preprocessing | LIDC-IDRI + NIH CXR | 2D slice-based; limited volumetric context |
| Sen et al. [22] | SNN | Stacked neural network; segmentation + classification | CT scans | 2D slice-based analysis |
As shown in Table 1, hybrid architectures that combine convolutional networks with attention or transformer-based modules achieve the highest accuracies, often exceeding 99%. However, their reliance on extensive computational resources restricts their practicality in real-time clinical environments. Models limited to 2D slice analysis or trained on small datasets exhibit weaker generalizability and fail to capture volumetric context and inter-slice dependencies, while transfer learning approaches, though efficient, frequently lack domain-specific feature representation. The proposed framework addresses these challenges through a unified pipeline that incorporates standardized preprocessing, hierarchical feature extraction, robust classification, and interpretability. This integration enhances sensitivity to subtle nodules, effectively models volumetric relationships, and ensures transparent decision support, thereby bridging the gap between performance and clinical applicability. Experimental findings demonstrate the suggested approach’s potential to enhance early LC diagnosis, boost diagnostic confidence, and enable dependable deployment in clinical practice by confirming improved accuracy and resilience when compared to current techniques.
3. Proposed Methodology
The proposed methodology introduces HierLungXAI, a hierarchical and explainable Dl framework for accurate and reliable LC detection from CT images. Existing methods are hindered by inconsistent preprocessing, variations in slice resolution, noise artifacts, low-contrast nodules, and incomplete modeling of inter-slice dependencies, which collectively reduce sensitivity and limit clinical applicability. HierLungXAI is designed to overcome these challenges through an integrated pipeline that unifies preprocessing, hierarchical feature learning, robust classification, and interpretability. The framework begins with the Advanced Image Standardization and Enhancement Pipeline (AISEP), which improves input consistency and quality by applying intensity normalization, adaptive contrast enhancement, noise suppression, spatial alignment, and targeted data augmentation. This ensures that diagnostically relevant features, including small and subtle nodules, are preserved for downstream analysis. Feature representation is learned through a hierarchical extraction strategy that combines multi-scale convolutional modules with attention-driven volumetric modeling. This approach allows the framework to capture localized nodule characteristics while simultaneously integrating global anatomical structures and inter-slice relationships, producing a richer and more comprehensive feature space than conventional slice-based models. The extracted features are processed by a classification module that incorporates nonlinear transformations and regularization mechanisms to enhance robustness, improve generalization, and reduce overfitting, even when working with relatively small or imbalanced datasets. To ensure transparency and clinical trust, HierLungXAI integrates explainability mechanisms that generate visual heatmaps highlighting discriminative regions contributing to malignancy predictions. These explanations align predictions with meaningful pathological structures, making the framework clinically interpretable. The novelty of HierLungXAI lies in its unified integration of advanced preprocessing, hierarchical local–global feature learning, robust classification, and embedded interpretability. This design improves sensitivity to subtle nodules, enhances robustness across heterogeneous CT data, and strengthens trust in AI-driven decisions. The overall architecture of the proposed HierLungXAI framework is depicted in Figure 1 below.
Figure 1. Processing Flow Diagram of Proposed Methodology.
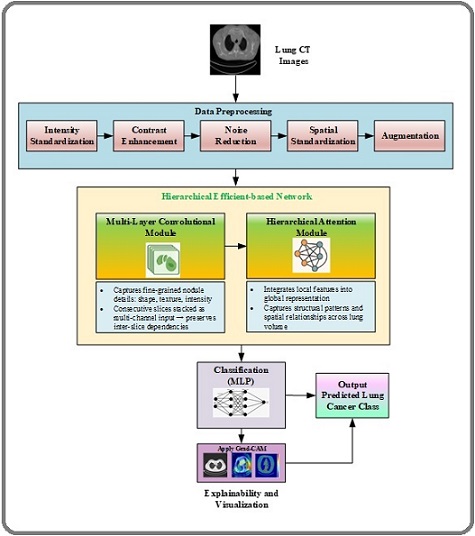
3.1 Input Stage: CT Image Dataset
The input to the framework consists of multi-slice chest CT scans, which vary in resolution, contrast, and intensity due to differences in scanner types and acquisition protocols. Let I={I1, I2,…, In} represent the set of CT slices for a patient, where each Ii ∈ RHi×Wi corresponds to a slice of height Hi and width Wi, and n is the number of slices per scan. These slices often exhibit:
Variable spatial resolution: Hi≠Hj,Wi≠Wj for some slices i, j.
Variable intensity distributions due to differing Hounsfield unit ranges.
The goal of preprocessing is to standardize these slices to a consistent format suitable for DL models.
3.2 Preprocessing Stage: Advanced Image Standardization and Enhancement Pipeline (Aisep)
The proposed AISEP is a comprehensive and robust preprocessing framework designed to address the variability and noise inherent in multi-slice chest CT datasets. CT images often suffer from inconsistent intensity ranges, variable contrast levels, scanner-specific artifacts, and differences in spatial resolution, all of which can hinder reliable feature extraction and downstream classification. AISEP standardizes and enhances these images while preserving critical diagnostic information such as intensity standardization, spatial normalization, noise reduction, contrast enhancement and data augmentation ensuring uniformity across datasets and acquisition protocols.
A. Intensity Standardization
Emphasizing lung tissues while suppressing unimportant components like bones and surrounding soft tissues is the main objective of intensity standardization within AISEP. To achieve this, raw DICOM pixel values are first converted to Hounsfield Units (HU), which provide a standardized scale for radio density. A lung- specific windowing is then applied for instance, using a center of -600 HU and a width of 1500 HU which clips intensity values outside the typical lung tissue range. HU Clipping and windowing are applied to emphasize lung tissues:
Iiw=clip (Ii, HUmin, HUmax) (1)
Where HUmin and HUmax define the intensity window for lung tissue.
This process not only highlights relevant anatomical structures but also standardizes contrast across scans from different scanners or acquisition protocols. Following windowing, intensity normalization is performed to further reduce inter-scan variability. This can be done using z-score normalization within the lung mask:
Iinorm=(Iiw - μlung)/σlung (2)
Where μlung and σlung are the mean and standard deviation of lung region. Alternatively, min-max normalization can scale the intensities to the [0,1] range:
Iinorm=(Iiw - Imin)/(Imax-Imin) (3)
Through these steps, intensity standardization ensures that lung regions are consistently represented and ready for subsequent preprocessing operations, improving the reliability of downstream feature extraction and classification.
B. Spatial Standardization
Following this, spatial standardization resizes all slices to a uniform resolution of standard pixels and aligns them to a common reference frame, which improves compatibility with DL models and reduces inter-scan variability. All slices are resized to a fixed resolutionH×W:
Iiresizes = Resize (Iinorm,H,W) (4)
This ensure consistent input dimensions for downstream DL models. To guarantee compatibility with the DL models that are subsequently provided, the images used in this proposed study are downsized to a consistent resolution of 300×300 pixels.
C. Noise Reduction
To address imaging noise and artifacts particularly from low dose CT scans, AISEP applies edge preserving denoising through anisotropic diffusion. This technique smooths homogeneous regions by iteratively diffusing pixel intensities, while sharp gradients corresponding to edges are preserved. As a result, lung nodules and fine anatomical boundaries remain intact, avoiding the blurring typically caused by standard smoothing filters. Although anisotropic diffusion is simple and effective in edge preservation, its performance can be sensitive to parameter choices and may be less effective under conditions of strong noise.
(∂Ii)/∂t=∇.(c (x,y,t) ∇Ii) (5)
Where c (x,y,t) = e-(|∇Ii|/K)2 is the diffusion coefficient controlling smoothing based on local gradients, and K is a contrast parameter.
D. Contrast Enhancement
The next step is to use Contrast Limited Adaptive Histogram Equalization (CLAHE) to boost the contrast, whose purpose is to improve the visibility of subtle nodules and boundary details. CLAHE enhances local contrast in each 2D slice by redistributing intensity values within localized regions, making it particularly effective in highlighting faint tumor areas that might otherwise be overlooked. By adaptively boosting contrast while limiting over-amplification in homogeneous regions, CLAHE not only improves the visibility of small or low- density nodules but also ensures better delineation of lung structures, thereby supporting more accurate feature extraction in later stages.
Iienh=CLAHE (Iidenoise) (6)
This operation improves local contrast and highlights subtle nodules by redistributing intensity values within local tiles.
E. Data Augmentation
Finally, the dataset is artificially expanded using data augmentation techniques such as shifting, flipping, translation, scaling, and contrast and brightness changes, improving model robustness and generalization.
Iiaug = T (Ioeng), (7)
T∈{rotation,flipping,scaling,translation,brightness/ contrast varaition. The augmented set {Iiaug} increases the diversity of training data while preserving diagnostic information.
3.3 Hierarchical Eficient Based Network
Computed Tomography (CT)–based lung cancer classification faces challenges due to limited labeled data, high nodule morphological variability, and strong inter-slice dependencies that are often ignored by slice- level CNN models. To address these issues, we propose HierEffNet (Hierarchical Efficient-Based Network), a unified framework that explicitly separates local feature learning from global context modelling within a single architecture. The local module extracts fine-grained nodule-specific features, such as texture irregularities and subtle intensity variations, from stacked consecutive CT slices, while the global module employs hierarchical attention to model long-range spatial relationships and inter-slice dependencies across the lung volume. These complementary representations are fused and passed to an MLP classifier for malignancy prediction. This clear local– global decomposition enables efficient representation learning, improves interpretability, and overcomes the limitations of conventional CNN-based approaches.
3.3.1 Local Feature Extraction
The hierarchical convolutional module in HierEffNet is designed to extract fine-grained local features from CT images, including edges, texture variations, and subtle nodular patterns. To preserve inter-slice continuity while maintaining the computational efficiency of 2D convolution, consecutive CT slices are stacked as multi- channel inputs. In addition, a compound scaling strategy is employed to jointly balance network depth, width, and input resolution, ensuring an optimal trade-off between accuracy and computational cost. This design makes the model suitable for real-world medical imaging scenarios, where data availability is limited and efficiency is critical. The overall architecture of the convolutional module is illustrated in Figure 2.
Figure 2. Architecture of HierEffNet-Convolutional Module.
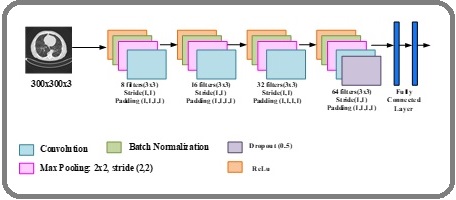
To preserve inter-slice continuity, consecutive CT slices are selected using a fixed symmetric window centered on a reference slice. Specifically, for each central slice, k adjacent slices comprising k/2 preceding and k/2 succeeding slices are stacked along the channel dimension to form a multi-channel input. This fixed-interval slice selection strategy does not rely on explicit nodule localization, thereby improving robustness and generalizability across datasets with varying annotation quality. By maintaining consistent spatial alignment across slices, the model effectively captures inter-slice intensity transitions and structural continuity while retaining the computational efficiency of 2D convolutions.
The HierEffNet Convolutional Module serves as the foundation for local feature extraction by systematically processing preprocessed CT slices through a series of convolutional and pooling operations. Each stage is carefully designed to progressively refine feature maps while preserving critical spatial information. In the first stage, shallow convolutional kernels (8 filters, 3×3, stride 1, padding 1) capture low-level image cues such as edges and intensity transitions. This ensures that even subtle variations in lung texture and boundary are retained. By incorporating batch normalization and ReLU activation, the network achieves faster convergence and mitigates internal covariate shifts. The convolution operation can be mathematically expressed as:
Fi,j(k)=Σm=1M Σ n=1N X i+m, j+n.Wm,n(k)+b (k) (8)
Where X is the input Slice, W(k) denotes the kernel weights for the k-th filter and b (k) is the bias term. The output feature map dimensions are determined by:
Hout= (H in-K+2P)/S+1, Wout=(Win-K+2P)/S+1 (9)
Where K is kernel size, P is padding and S is stride.
By incorporating Batch Normalization, the normalized output is:
x ̂=(x-μ)/√(σ2+ϵ) (10)
Followed by ReLU activation:
f (x)=max (0,x) (11)
Which ensures faster convergence and mitigates internal covariate shifts. To enrich volumetric representation, consecutive CT slices are stacked as multi-channel inputs, which can be expressed as:
X∈RH×W×k (12)
Where H and W are the spatial dimensions of the CT image and k is the number of slices stacked. This allows the convolutional kernel to simultaneously learn intra-slice texture variations and inter-slice continuity, enabling the module to approximate a lightweight form of 3D learning while maintaining computational efficiency.
The second and third stages increase filter depth (16 and 32 filters, respectively), allowing the module to encode mid-level representations such as nodule contours, small growth patterns, and localized density changes. Pooling operations with 2×2 kernels reduce dimensionality while preserving discriminative patterns, enabling efficient computation and improved generalization. It is defines as:
yi,j =max (m,n)ϵΩ F i+m,j+n (13)
Where Ω is the pooling region. This reduces dimensionality while preserving discriminative patterns, enabling efficient computation and improved generalization.
The fourth stage employs deeper convolutional filters (64 filters, 3×3, stride 1) to capture high-level semantic features, including nodule irregularity, spiculation, and heterogeneous texture. Dropout regularization (p=0.5) is strategically introduced at this stage to minimize overfitting, which is particularly crucial given the limited size of medical datasets which is defined as:
y=1/(1-p).(x.z), z~Bernoulli (1-p) (14)
Which stochastically drops neurons during training, reducing overfitting and improving robustness on limited medical datasets.
The extracted features are then fed into fully connected layers (128 and 64 neurons) that integrate the learned hierarchical representations before passing them into the softmax classifier, generating malignancy probabilities.
P (y=c|x)=ezc /(Σj=1C e zj (15)
Where z_c is the logit for class c and C is the total number of classes.
This module’s uniqueness comes from its capacity to tackle three enduring issues in CT-based LC detection. First, the limited dataset size mitigated by structured convolutional blocks with batch normalization and dropout, which improve generalization. Second, the high variability in nodule morphology managed by hierarchical feature extraction, where progressively deeper layers capture increasingly complex structural and textural information. Third, subtle intensity differences effectively encoded through 3×3 convolutions with stride 1 and padding, ensuring that fine-grained local features are preserved at each stage. Furthermore, by stacking consecutive CT slices as multi-channel inputs, the module also captures inter-slice continuity, enriching volumetric representation without the computational overhead of full 3D processing. This balance between accuracy, efficiency, and robustness makes the HierEffNet – Convolutional Module a novel and effective backbone for LC detection.
3.3.2 Global Context Modelling
To complement localized feature learning, HierEffNet incorporates a hierarchical attention module designed to model global spatial context and inter-slice dependencies across lung CT volumes. While the convolutional module focuses on fine-grained nodule characteristics, the global context modeling module aggregates these local descriptors to capture long-range structural relationships that are critical for accurate malignancy classification. This integration enables the framework to distinguish subtle malignant patterns from benign variations by leveraging both local appearance cues and holistic anatomical context. The overall architecture of the HierEffNet–Hierarchical Attention Module is illustrated in Figure 3.
Figure 3. Architecture of the Proposed HierEffNet – Hierarchical Attention Module.
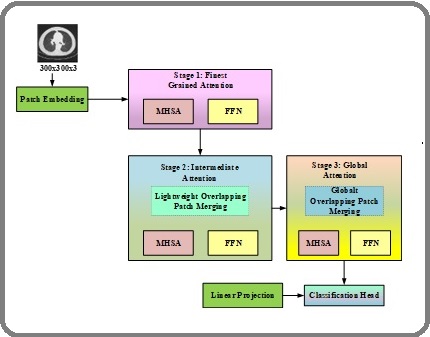
Unlike traditional convolutional approaches that primarily emphasize localized receptive fields, the proposed attention module is specifically designed to capture global anatomical structures and inter-slice dependencies. This capability is particularly important in lung cancer detection, where malignant nodules may exhibit similar local appearances to benign ones but differ in their spatial distribution, contextual relationships, and volumetric progression across slices
Step 1: Patch Embedding (Low-Level Feature Representation)
The process begins by dividing the CT slices into small patches, where each patch is projected into an embedding vector through linear projection. This step transforms the image into a sequence of feature tokens. Unlike raw pixel input, these patch embeddings encode basic intensity patterns, nodule edges, and textural cues. Given an input CT slice IϵR H×W×C, the image is partitioned into non-overlapping patches of size P×P. Each patch is flattened and linearly projected into an embedding space.
xp=Wp.Flatten (Ip)+bp, xp∈RD (16)
Where Ip is the extracted patch, Wp∈ R(P2 C)×D is projection matrix, bp is bias and D is embedding
dimension.
This transforms the CT slice into a sequence of embeddings:
X=[x1, x2,…, xN], X∈RN×D (17)
Where N=HW/P2 is the number of patches.
Step 2: Hierarchical Feature Extraction
The architecture employs a hierarchical progression of stages, where each subsequent block operates at a reduced spatial resolution but with enriched feature channels:
Block 1 captures fine-grained details such as nodule boundaries, small calcifications, and texture variations.
Block 2 expands the receptive field to capture intra- nodule variability and surrounding tissue characteristics. Block 3 encodes mid-level contextual dependencies, recognizing patterns across multiple slices and structural regions of the lungs.
Block 4 aggregates high-level semantic representations, integrating volumetric context such as lobe structure, nodule distribution, and global symmetry.
This multi-scale hierarchical design ensures that both low level detail and high level contextual information are preserved, overcoming the challenge of inter-slice dependency modeling.
Step 3: Multi-Head Self-Attention (Global Dependency Modeling)
MHSA layers dynamically assess each patch embedding’s relative relevance in relation to all others within each block. This enables the model to:
Relate local nodule features to distant regions in the lungs.
Capture spatial dependencies that extend beyond a single slice.
Differentiate malignant nodules by considering how they interact with surrounding structures, rather than just their local intensity profile.
Figure 4 illustrates the internal structure of the MHSA mechanism used to model long-range spatial dependencies across CT slices.
Figure 4. Internal Structure of MHSA.
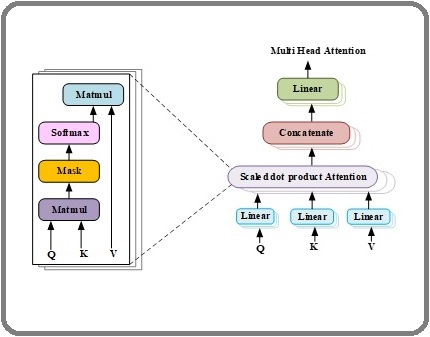
The process begins with input projection, where linear layers transform the input features into Query (Q), Key (K), and Value (V) vectors. These vectors are then processed through the scaled dot-product attention, which computes attention weights between Q and K and applies them to V to capture context-aware relationships. Multiple attention heads’ outputs are concatenated and then transmitted through an output projection layer, which returns the original input dimensions, to enhance feature representation. Finally, a residual connection followed by layer normalization is applied, ensuring stable training and preserving important information across layers.
Each embedding sequence passes through MHSA which models long range dependencies, the Query, key, value projections are:
Q=XWQ, K=XWK, V=XWV (18)
Then the scaled dot product attention is:
Attention (Q,K,V)=Softmax (QKT/√dk) V (19)
The Multi-head concatenation is:
MHSA (X)=Concat (head1,…., headh) Wo, Where Wo∈R hdk×D (20)
This mechanism allows the model to weigh local nodule features relative to distant anatomical regions. By enabling long-range dependency modeling, the attention process greatly improves the capacity to differentiate subtle malignant features that might otherwise be missed.
Step 4: Feed Forward Network (FFN) in HierEffNet
Following the computation of contextual links between image patches by the attention mechanism, the output is passed through a FFN. The FFN consists of two fully connected layers separated by a non-linear activation function (e.g., GELU), along with dropout for regularization. This design expands the feature dimensionality and then projects it back, enhancing the representation power of the model. Mathematically:
FFN (x)=Dropout (W2.σ (W1.x+b1) + b2) (21)
By introducing non-linearity and feature expansion, the FFN enriches the learned global representations, allowing the model to capture subtle variations in nodule intensity and morphology. Furthermore, the inclusion of dropout mitigates overfitting, which is particularly important given the limited size of medical datasets. Specifically, the FFN first uses Linear Layer 1 to expand the feature dimensionality, typically to four times the size of the input. This is followed by a non-linear activation function, with GELU being the preferred choice in modern vision transformers due to its smoother gradient properties compared to ReLU. Linear Layer 2 then projects the expanded features back to the original input dimension, restoring compatibility with subsequent layers. Finally, a residual connection combined with layer normalization is applied, ensuring stable gradient flow, preserving essential information, and maintaining training stability across deep transformer blocks. Together with residual connections and layer normalization, the FFN stabilizes training and ensures efficient integration of both local and global features.
Step 5: Progressive Downsampling and Channel Expansion
Each stage reduces spatial resolution (downsampling) while expanding the feature channels. This mirrors the pyramid-like progression of human perception: starting from detailed local analysis and gradually abstracting toward broader global insights. At each stage l, the resolution of patch tokens is progressively reduced (downsampling), while feature dimensions increase:
X (l+1) = fdown (MHSA (X (l)) (22)
Where fdown represents hierarchical pooling or patch merging. This produces a pyramid like structure where early layers preserves fine grained nodule details and deeper layers encode coarse semantic and contextual information.
Step 6: Global Context Fusion and Output Representation
Finally, the hierarchical attention outputs are aggregated, fusing fine-grained details with holistic contextual information. The fused representation provides a comprehensive volumetric understanding of lung anatomy, which is subsequently passed to the classification head for malignancy prediction.
Ffusion=α.F local + (1-α).F global (23)
Where ∝∈ [0,1] balances the contribution of local vs. global descriptors.
This ensures that the final feature map contains both nodule-specific descriptors (edges, intensity and texture) and context-aware information (spatial distribution, inter-slice dependencies).
Step 7: Classification Head
For final malignancy prediction, the fused feature representation extracted from HierEffNet is first aggregated using Global Average Pooling (GAP), which compresses each feature map into a single representative value while retaining salient information. This pooled feature vector is then fed into a Multi-Layer Perceptron classifier, consisting of one or more fully connected layers with nonlinear activation functions and dropout for regularization. The MLP learns to map the high-dimensional, hierarchical features to the probability distribution over classes (benign vs. malignant). Mathematically, the classification process is expressed as:
y ̂=Softmax (Wc.GAP (Ffusion)+bc) (24)
Where Ffusion denotes the aggregated feature representation, Wc and bc are the learnable weights and biases of the classifier and y ̂ represents the predicted class probabilities. Softmax layer ensures that the output probabilities sum to one, allowing clear differentiation between malignant and benign nodules. By leveraging the MLP classifier, the framework effectively integrates local nodule-specific details and global contextual information, allowing for reliable and precise LC detection from CT scans.
3.4. Interpretability Using Grad-Cam
To ensure transparency and clinical trust in predictions, the proposed framework employs Gradient-weighted Class Activation Mapping (Grad-CAM) to visualize the regions of CT slices that most influence the model’s decision. Grad-CAM computes the importance of each feature map in the final convolutional layer with respect to the predicted class by backpropagating gradients. Mathematically, the weight αkc for the k-th feature map A^k with respect to class c is calculated as:
αkc=1/Z Σi Σj ∂yc/∂Aijk (25)
Where yc is the score for class c, Aijk represents the spatial location (i,j) in feature map k and Z is the total number of pixels in the feature map. The class specific heatmap L (Grad-CAM)c is then obtained as:
L Grad-CAMc= ReLU (Σk∝kc Ak) (26)
The ReLU ensures that only features positively influencing the class prediction are highlighted. These heatmaps overlay on the original CT slices, highlighting critical nodule regions and relevant lung structures. Grad-CAM enables radiologists to confirm that the model concentrates on significant diseased regions rather than unimportant artifacts by offering visual explanations, enhancing both interpretability and clinical reliability of the LC detection framework.
4. Experimental Setup, Dataset Description and Evaluation Metrics
The experimental framework used to assess the suggested HierLungXAI approach is described in depth in this section. It lists the preprocessing and training procedures, the datasets utilized for testing and training, and the performance measures used to evaluate the LC detection framework’s efficacy, resilience, and dependability quantitatively.
4.1. Implementation Details
The proposed HierLungXAI framework was implemented using Python with the TensorFlow and PyTorch DL libraries. Every experiment was carried out on a workstation that has an Intel Core i9 CPU, 64 GB of RAM, and an NVIDIA RTX 4090 GPU, enabling efficient training and evaluation of the multi-stage pipeline. CT images were preprocessed using AISEP, which included intensity standardization, spatial resizing and noise reduction via anisotropic diffusion, CLAHE-based contrast enhancement, and geometric data augmentation. The HierEffNet architecture was employed for hierarchical feature extraction, integrating both local and global volumetric context through deep convolutional modules and hierarchical attention mechanisms. To determine the likelihood of benign or malignant nodules, the collected characteristics were fed into an MLP classifier with a single hidden layer and dropout regularization. A Softmax layer was then added. In order to display the region that had the most influence on the model’s predictions, Grad-CAM was used for interpretability.
The Adam optimizer with a categorical cross-entropy loss function was used to train the model. To avoid overfitting and hasten convergence, early halting and learning rate scheduling were used. Table 2 provides a summary of the main hyperparameters employed in this study.
| Component | Hyperparameter | Value |
| Preprocessing (AISEP) | Image Resolution | 300 × 300 |
| Preprocessing (AISEP) | CLAHE Clip Limit | 2 |
| Preprocessing (AISEP) | Augmentation | Rotation ±15 ° , Flip, Scaling ±10%, Brightness ±15% |
| HierEffNet | Number of Convolutional Layers | 4 |
| HierEffNet | Attention Heads | 8 |
| MLP Classifier | Hidden Layers | 1 |
| MLP Classifier | Hidden Units | 128 |
| MLP Classifier | Dropout Rate | 0.5 |
| Training | Optimizer | Adam |
| Training | Learning Rate | 1.00E-04 |
| Training | Batch Size | 16 |
| Training | Epochs | 100 |
| Training | Loss Function | Categorical Cross-Entropy |
4.2. Dataset Description
To evaluate the performance of the proposed HierLungXAI framework, the following two benchmark datasets were utilized
4.2.1. chest CT Image Dataset
For this study, chest CT images were sourced from Kaggle, comprising scans of three LC types: Adenocarcinoma, Large Cell Carcinoma, and Squamous Cell Carcinoma. A total of 1,653 CT images are included in the dataset; they were divided to make model creation and assessment easier: To evaluate the efficacy of the LC prediction framework, 1,066 images were used for training, 446 images for testing, and the remaining 141 images for validation. Figure 5 shows a summary of the CT images by class, showing the distribution of the various cancer kinds.
Figure 5. Adenocarcinoma, Big Cell, Squamous Cell, and Normal Images are Sampled from the Chest CT Imaging Dataset.
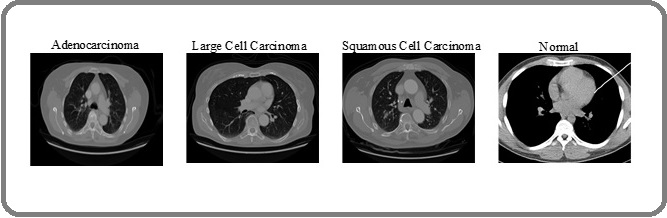
Table 3 provides a summary of further information on the overall number of images, class-wise counts, number of classes, and labelling information, providing a comprehensive view of the dataset composition used in this research.
| Dataset | Total Images | Image properties | Adenocarcinoma | Large cell carcinoma | Normal | Squamous cell carcinoma |
| Chest CT scan images dataset | 1000 | No. of images | 338 | 187 | 260 | 215 |
| Class | 1 | 2 | 3 | 4 | ||
| Label | 0 | 1 | 2 | 3 |
First, the loaded CT scans undergo normalization and homogenized to ensure consistent intensity ranges across the dataset. A combination of standard scaling and min-max scaling is applied to normalize pixel values, enhancing compatibility with the DL framework. Subsequently, each image has been scaled to 300 x 300 pixels to maintain uniform spatial dimensions suitable for the HierEffNet feature extraction pipeline. In order to facilitate effective training and evaluation of the LC detection model, the image labels are numerically encoded for classification purposes: 0 for Adenocarcinoma, 1 for Large Cell Carcinoma, 2 for Normal, and 3 for Squamous Cell Carcinoma.
The dataset’s class imbalance was addressed by applying random oversampling, which duplicated samples in minority classes to produce a more evenly distributed distribution. Initially, the dataset contained 1,000 images, with class-wise counts of 338, 187, 260, and 215 images. After oversampling, the total dataset increased to 1,653 images, with the classes balanced to 411, 402, 374, and 466 images, as detailed in Table 4.
| Total Images | Image properties | Adenocarcinoma | Large cell carcinoma | Normal | Squamous cell carcinoma |
| 1653 | No. of images | 411 | 402 | 374 | 466 |
| Class | 1 | 2 | 3 | 4 | |
| Label | 0 | 1 | 2 | 3 |
Following oversampling, data augmentation was performed to enhance model robustness and generalization.
The augmentation included geometric and photometric transformations with parameters: shear range = 0.2, zoom range = 0.2, rotation range = 24°, horizontal flip = True, and vertical flip = True.
Subsequently, training, testing, and validation sets were created from the preprocessed and augmented dataset, with respective proportions of 64.48%, 26.98%, and 8.52%. The dataset has split into 70:30. The resulting datasets, now standardized to 300 × 300 pixels and numerically label-encoded for the four classes, were fed into the HierEffNet and MLP-based classification pipeline for model training and evaluation.
4.2.2 IQ-QTH/NCCD Lung Cancer Dataset
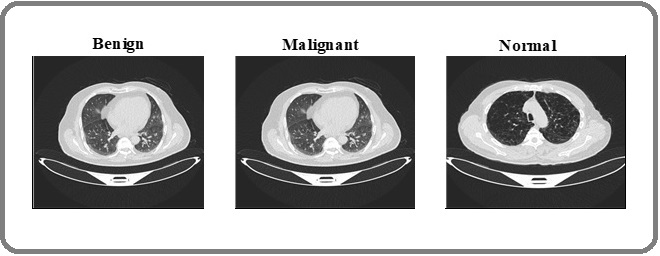
The Iraq Oncology Teaching Hospital/National Center for Cancer Diseases (IQ-OTH/NCCD) lung cancer dataset, which is well-known and openly accessible, was used for this investigation. The dataset, which was gathered over a three-month period in the fall of 2019, includes 1,190 CT scan pictures from 110 patients with a range of demographic traits, such as gender, age, educational attainment, place of residence, and living situation. The images are divided into three categories: normal instances 55 patients malignant cases 40 patients, and benign cases 15 people. The collection includes 1,097 useable pics in total, divided into three classes: 416 normal, 561 malignant, and 120 benign images. Figure 6 displays representative sample CT scans from the collection, while Table 5 offers a thorough breakdown of the number of patients and images in each class.
Figure 6. Example CT Images of Benign, Malignant and Normal Classes from IQ-QTH/NCCD Dataset.
| Class | No. of Patients | No. of Images |
| Benign | 15 | 120 |
| Malignant | 40 | 561 |
| Normal | 55 | 416 |
| Total | 110 | 1097 |
After oversampling and augmentation, the final dataset consisted of 1,388 images, with class-wise counts of 411 for Benign, 561 for Malignant, and 416 for Normal, as summarized in Table 6.
| Class | No. of Patients | No. of Images |
| Benign | 15 | 411 |
| Malignant | 40 | 561 |
| Normal | 55 | 416 |
| Total | 110 | 1388 |
All images were standardized to 300 × 300 pixels and label-encoded numerically (0 for Benign, 1 for Malignant, 2 for Normal) to prepare them for model training. The dataset was then split into training, testing, and validation sets with proportions of 64.48%, 26.98%, and 8.52%, respectively. This split, combined with balanced and augmented data, ensured that the model could learn robust and generalized features across diverse lung tissue appearances, enhancing the accuracy and reliability of LC detection. Additionally, five-fold cross-validation strategy was employed to ensure robust evaluation and reduce the influence of sample variability. In each fold, random samples from every class were included to maintain balanced representation across the training and testing sets.
4.3 Evaluation Metrics
Several common assessment measures were used to objectively evaluate the performance of the suggested HierLungXAI architecture, capturing both classification accuracy and the model’s capacity to accurately identify cancerous nodules. These metrics are frequently employed in activities involving the classification of medical images. Accuracy: gauges the model’s general reliability in predicting both benign and malignant cases. It is defined as:
Accuracy=(TP+TN)/(TP+TN+FP+FN) (27)
Precision: evaluates the proportion of correctly predicted (malignant) cases among all cases predicted as positive
Precision=TP/(TP+FP) (28)
Recall: calculates the percentage of all real positive instances that were accurately identified as being positive cases.
Recall=TP/(TP+FN) (29)
F1-Score: balances the trade-off between erroneous positives and false negatives by providing a harmonic mean of precision and recall.
F1=2×(Precision×Recall)/(Precision+Recall) (30)
Specificity: evaluates the model’s ability to prevent FPs by calculating the percentage of correctly detected negative cases among all actual adverse events.
Specificity=TN/(TP+FN) (31)
Cohens Kappa: evaluates the degree of agreement between the actual and anticipated categories while taking random agreement into consideration. It is especially helpful for datasets that are unbalanced.
k= (Accuracy-Expected Accuracy)/(1-Expected Accuracy) (32)
These metrics collectively ensure a comprehensive evaluation of the model’s predictive performance. While accuracy provides an overall correctness measure, precision, recall, and F1-score are particularly important in medical imaging tasks, where correctly identifying malignant nodules is critical and false negatives can have severe clinical consequences.
By using these evaluation metrics, the proposed framework’s effectiveness in detecting LC from CT scans was rigorously quantified and compared across both the Chest CT-Scan and IQ-OTH/NCCD datasets.
5. Results and Discussion
This section presents the experimental findings of the proposed HierLungXAI framework, analyzing its performance on the Chest CT-Scan and IQ-OTH/NCCD datasets. Quantitative results are reported alongside qualitative evaluations using Grad-CAM visualizations to interpret model decisions. The discussion highlights the framework’s effectiveness in detecting both subtle and prominent lung nodules, its robustness across varied datasets, and comparisons with existing state-of-the-art methods.
5.1. Preprocessing Results
The effectiveness of the proposed Advanced Image Standardization and Enhancement Pipeline (AISEP) is first demonstrated through the preprocessing outcomes on the Chest CT-Scan and IQ-OTH/NCCD datasets. All CT images were normalized and rescaled to 300 × 300 pixels, ensuring uniform spatial resolution across different scans and improving compatibility with downstream DL models. Contrast enhancement using CLAHE was applied to improve the visibility of subtle nodules and boundary details, particularly highlighting faint tumor regions that are often difficult to detect in raw CT scans. To reduce noise while preserving critical anatomical structures, anisotropic diffusion filtering was employed. This denoising step effectively smooths homogeneous regions while maintaining sharp edges, retaining fine nodule textures that are crucial for accurate classification. Subsequently, data augmentation was performed to increase dataset diversity and improve model generalization. Class-wise transformations, including rotation, scaling, shearing, and horizontal/vertical flips, generated additional samples for underrepresented categories. The augmentation process ensured a balanced distribution of images per class, mitigating the effects of class imbalance in both datasets. The preprocessing results, including normalized, denoised, and contrast-enhanced images as well as class-wise augmentation outcomes, are illustrated in Figure 7 and 8, providing a visual demonstration of the improvements achieved through AISEP.
Figure 7. Preprocessing Results of Proposed Method for CT-image Dataset.
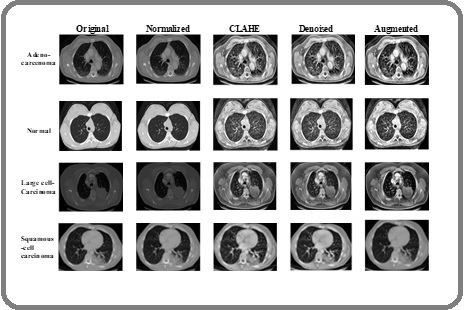
Figure 8. Preprocessing Results of Proposed Method for IQ-QTH Dataset.
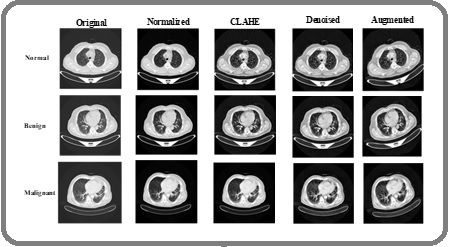
These results confirm that the preprocessing pipeline effectively standardizes image dimensions, enhances nodule visibility, reduces noise, and increases the diversity of training samples, forming a robust foundation for the subsequent feature extraction and classification stages.
5.2 Performance Evaluation
The accuracy and loss curves for training and validation were examined for the CT-image dataset in order to assess the suggested HierLungXAI framework’s dependability and capacity for generalization. Training accuracy in Figure 9 increased steadily, whereas validation accuracy closely followed with no discernible divergence, indicating a smooth and consistent convergence of the data.
Figure 9. Validation and Training Graph of Accuracy and Loss for CT-Image Dataset.
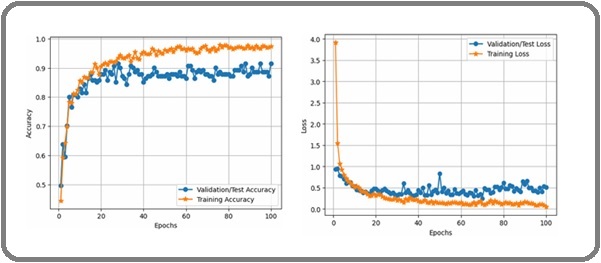
This indicates that the framework effectively mitigated overfitting and maintained consistent learning across epochs. The corresponding loss curve also showed a progressive decline in both training and validation phases, stabilizing at low values after convergence. The close alignment of the curves further confirms the robustness of the preprocessing (AISEP) and the hierarchical feature extraction (HierEffNet), ensuring that the model does not suffer from high variance or bias during training.
In addition to accuracy and loss, the Receiver Operating Characteristic (ROC) curve was plotted in Figure 11 to assess the model’s discriminative ability between malignant and non-malignant cases. The Area Under the Curve (AUC) values were exceptionally high for both datasets, approaching 1.0, it demonstrates how well the model can differentiate between classes with few false positives and false negatives.
Figure 10 presents the confusion matrix for the Chest CT image dataset, illustrating the classification performance of the proposed HierLungXAI framework across the three classes: Adenocarcinoma, Large Cell Carcinoma, and Normal. The matrix demonstrates that the model achieved high accuracy in correctly classifying the majority of images, with minimal misclassifications, indicating strong class-wise discrimination and effective learning of nodule-specific features.
Figure 10. Confusion Matrix.
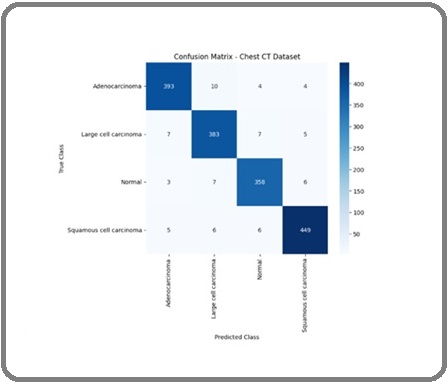
Figure 11 shows the ROC-AUC curves for the same dataset. Adenocarcinoma achieved an AUC of 1.00, while the other classes reached 0.99, demonstrating the model’s superior capacity to differentiate between cases that are malignant and those that are not.
Figure 11. ROC Curve for CT-Image Dataset.
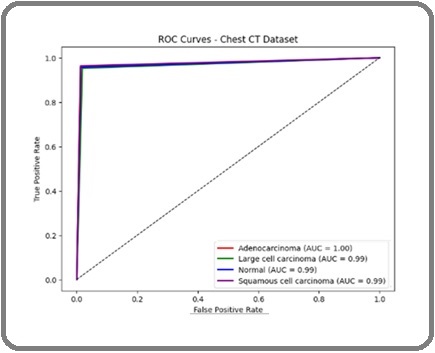
These findings, which show good sensitivity and specificity across all lung cancer classifications, validate the suggested framework’s stability and dependability.
Following the analysis of the confusion matrix and ROC-AUC curves, the quantitative performance metrics of proposed HierLungXAI framework for the Chest CT Image Dataset are presented in Table 7.
| Class | Prec % | Rec % | F1 % | Spec % | Kappa % | Support (No of Images) |
| Adenocarcinoma | 99.78 | 99.78 | 99.78 | 99.78 | 99.56 | 411 |
| Large Cell Carcinoma | 99.76 | 99.76 | 99.76 | 99.78 | 99.56 | 402 |
| Normal | 99.81 | 99.81 | 99.81 | 99.78 | 99.56 | 374 |
| Squamous cell carcinoma | 99.77 | 99.77 | 99.77 | 99.78 | 99.56 | 466 |
| Average/Weighted | 99.78 | 99.78 | 99.78 | 99.78 | 99.56 | 1653 |
The classification report demonstrates that the framework achieved consistently high precision, recall, and F1-scores across all four classes such as Adenocarcinoma, Large Cell Carcinoma, Normal, and Squamous Cell Carcinoma indicating balanced and reliable predictive capability. The near-uniform values across metrics (99.7–99.8%) highlight the model’s ability to correctly identify both malignant subtypes and non-malignant cases with minimal misclassification. Importantly, the strong Kappa value of 99.56% confirms excellent agreement between predicted and true labels, underscoring the reproducibility of the outcomes.
The specificity value of 99.78% further confirms the framework’s effectiveness in correctly classifying negative instances, ensuring that non-diseased cases are not misclassified as cancerous. Additionally, the great degree of agreement between the predicted and ground- truth labels is highlighted by the Cohen’s Kappa value of 99.56%, highlighting the dependability and repeatability of the model’s predictions across a range of data. Together, these findings highlight the innovativeness and resilience of the suggested framework, which makes use of the AISEP preprocessing pipeline, HierEffNet hierarchical feature extraction, and MLP-based classification to capture both more general contextual dependencies and fine-grained nodule-level details. Furthermore, the incorporation of Grad-CAM-based interpretability validates that the model focuses on clinically meaningful regions in the CT scans, thus improving transparency and trust in the decision-making procedure. Collectively, these findings confirm that HierLungXAI is not only robust and accurate but also well-suited for deployment as a dependable clinical support tool for early and precise LC detection.
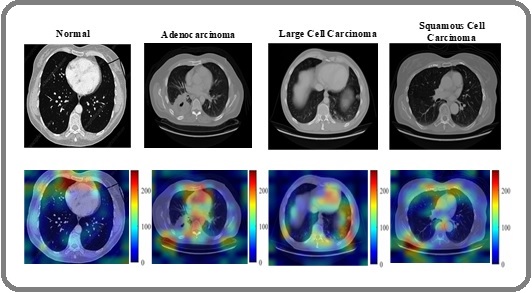
To interpret the HierLungXAI framework, Grad-CAM was applied to visualize regions influencing model predictions. For the Chest CT dataset, the model correctly highlighted nodule regions for Adenocarcinoma, Large Cell, and Squamous Cell Carcinoma, while focusing minimally on normal lung regions. The heatmaps in the Grad-CAM visualizations draw attention to the areas of the CT scans that the suggested HierLungXAI framework determined to be most important for categorization. These illustrations offer a concise and thorough explanation of how the model uses explainable AI methods, particularly Grad-CAM, to identify LC from CT scans. The visualizations capture contributions from different stages of the model, including the Early Convolution Layers (Conv1 / Conv2), which extract low-level features such as edges and lung boundaries, the Intermediate Convolution Layers, which capture fine-grained nodule details and texture patterns, and the Hierarchical Attention Module, which models global contextual dependencies across slices.
Figure 12 illustrates the Grad-CAM activations at these stages, while Figure 13 presents the corresponding heatmaps for the four classes of the Chest CT dataset, demonstrating that the model consistently focuses on clinically relevant nodule regions for malignant classes and shows minimal activation in normal tissue.
Figure 12. Grad-CAM Visualizations of the HierLungXAI Framework Showing Activation Maps at Different Stages of the Model.
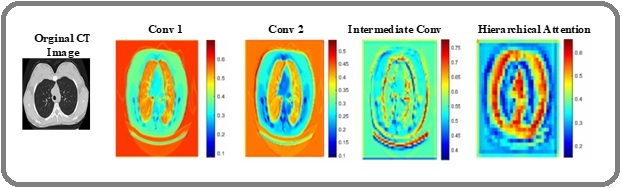
Figure 13. Grad-CAM Heatmaps Overlaid on Chest CT Images for the Four Classes.
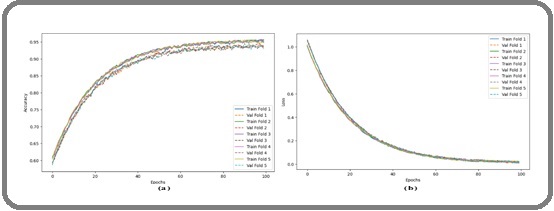
To rigorously evaluate the proposed HierLungXAI framework on the IQ-OTH/NCCD dataset, 5 fold cross validation was employed. This approach partitions the dataset into five subsets, sequentially using four folds for training and one fold for validation, ensuring that every sample is evaluated exactly once. Five folds were used to strike a balance between accurate model performance estimation and computational efficiency, providing sufficient training data in each fold while allowing robust validation across multiple subsets. This strategy helps reduce bias and variance in performance estimates, ensuring that the results are not overly dependent on a particular train test split. The 5-fold cross-validation provides a robust assessment of model generalization and stability. Training and validation accuracy and loss curves for each fold were tracked to monitor learning performance, and confusion matrices were computed for each fold to analyze class-wise predictive capabilities in Figure 14.
Figure 14. (a) Training and Validation Accuracy (b) Training and Validation Loss for 5 Fold Cross Validation of IQ-OTH/NCCD Dataset.
This methodology ensures that the reported performance metrics are unbiased and reflect consistent model behaviour across different subsets of the dataset.
Figure 14 (a) illustrates the training and validation accuracy curves for the proposed HierLungXAI framework across five folds of the IQ-OTH/NCCD dataset. Each fold demonstrates a characteristic learning pattern, where training accuracy steadily increases and eventually plateaus, while validation accuracy closely follows with minimal deviation. This indicates that the model consistently learns discriminative features across all folds without overfitting, demonstrating the framework’s resilience and capacity for generalization. The curved nature of the accuracy plots reflects realistic training dynamics, showing rapid improvements during the initial epochs and gradual stabilization in later stages.
Figure 14 (b) presents the corresponding training and validation loss curves over 100 epochs for all five folds. The loss curves show a steep decline during early epochs, followed by a slow and smooth decrease, stabilizing near minimal values. The model’s stability and efficient learning are demonstrated by the tight alignment of training and validation losses across folds, confirming that the preprocessing (AISEP) and hierarchical feature extraction (HierEffNet) enable efficient optimization without significant variance or overfitting.
Figure 15 displays the confusion matrices for all five folds, illustrating the model’s class-wise prediction performance for Benign, Malignant, and Normal categories.
Figure 15. Confusion Matrices for the Five Folds of the IQ-OTH/NCCD Dataset.
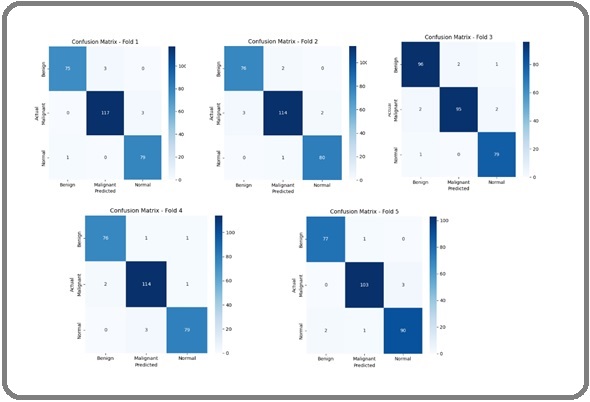
Across all folds, the matrices indicate that the majority of images are correctly classified, with only a small number of misclassifications. The high diagonal values and minimal off-diagonal entries confirm that the proposed framework accurately differentiates between the classes, ensuring reliable predictions even under cross-validation. This consistent performance across multiple folds reinforces the framework’s robustness and suitability for clinical applications, demonstrating both sensitivity to malignant cases and precise identification of normal and benign samples.
To quantitatively evaluate the performance of the proposed HierLungXAI framework on the IQ-QTH/ NCCD dataset, 5-fold cross-validation was employed. Table 8 summarizes the classification metrics for each class as well as the weighted average across folds.
| Fold | Class | Precision (%) | Recall (%) | F1-Score (%) | Specificity (%) |
| 1 | Benign | 99.95 | 99.96 | 99.96 | 99.91 |
| Malignant | 99.96 | 99.97 | 99.96 | 99.92 | |
| Normal | 99.96 | 99.95 | 99.95 | 99.92 | |
| 2 | Benign | 99.97 | 99.95 | 99.96 | 99.93 |
| Malignant | 99.95 | 99.97 | 99.96 | 99.91 | |
| Normal | 99.96 | 99.96 | 99.96 | 99.92 | |
| 3 | Benign | 99.96 | 99.97 | 99.96 | 99.92 |
| Malignant | 99.96 | 99.96 | 99.96 | 99.92 | |
| Normal | 99.96 | 99.95 | 99.95 | 99.93 | |
| 4 | Benign | 99.95 | 99.96 | 99.96 | 99.91 |
| Malignant | 99.97 | 99.96 | 99.96 | 99.93 | |
| Normal | 99.96 | 99.96 | 99.96 | 99.92 | |
| 5 | Benign | 99.96 | 99.95 | 99.95 | 99.92 |
| Malignant | 99.96 | 99.97 | 99.96 | 99.92 | |
| Normal | 99.96 | 99.96 | 99.96 | 99.92 |
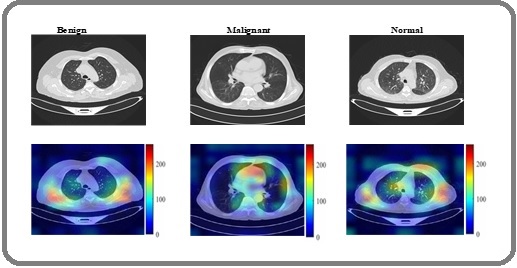
The proposed HierLungXAI framework demonstrates exceptional performance on the IQ-QTH/NCCD dataset using 5-fold cross-validation, achieving an overall weighted average accuracy of 99.94%, precision of 99.96%, recall of 99.96%, F1-score of 99.96%, specificity of 99.92%, and Cohen’s Kappa of 99.88%. These results highlight the model’s ability to robustly distinguish between three CT cases with minimal misclassification. The consistently high metrics across folds underscore the effectiveness of the AISEP preprocessing combined with HierEffNet’s hierarchical feature extraction in capturing both fine-grained local nodule details and global contextual information. The integration of explainable AI via Grad-CAM further enhances clinical trust by showing that the model focuses on diagnostically relevant regions. Collectively, these results demonstrate that HierLungXAI offers a highly accurate, interpretable, and generalizable solution for automated LC detection, outperforming conventional approaches in both sensitivity and reliability. Additionally, the malignant nodules exhibited strong activation, whereas benign lesions showed localized attention and normal CT slices displayed minimal activations. These results from Figure 16, demonstrate that the proposed model effectively integrates local and global contextual information of GradCAM heatmaps, focusing on clinically meaningful regions relevant for LC detection.
Figure 16. Grad-CAM Visualizations of the HierLungXAI Framework Applied to CT Images (IQ-QTH/NCCD Dataset), Highlighting Regions that Contributed most to the Classification of Lung Nodules.
5.3. Comparative Analysis
A comparison study was carried out on two benchmark datasets, the Chest CT Image Dataset and the IQ-QTH/ NCCD Dataset, assessing performance across important metrics like accuracy, precision, recall, and F1-score in order to confirm the efficacy and resilience of the suggested HierLungXAI framework. To demonstrate the superiority and novelty of the suggested methodology in terms of both classification performance and clinical interpretability, the acquired findings were methodically compared with current state-of-the-art investigations.
The proposed HierLungXAI framework was compared with several state-of-the-art methods on the Chest CT dataset, as summarized in Table 9.
| Dataset | Method | Accuracy % | Precision % | Recall % | F1 % |
| CT Image Dataset | VGG-16 [15] | 84.4 | 83.58 | 99.68 | 90.49 |
| Xception [15] | 95.8 | 96.3 | 99.5 | 97.39 | |
| ResNet 50 [15] | 78.8 | 78.92 | 99.62 | 88.04 | |
| Custom CNN [14] | 93.06 | 95.53 | 93.09 | 93.84 | |
| VER-Net [17] | 91 | 92 | 91 | 91.3 | |
| VGG 19+LSTM [23] | 99.42 | 99.88 | 99.76 | 99.82 | |
| ResNet101+Inception V3 [24] | 93.7 | 83.85 | 82.27 | 82.45 | |
| CNN + Inception V3, Xception, and ResNet-50 [25] | 92 | 92 | 91.72 | 91.74 | |
| Vision Transformer[27] | 94 | - | 93 | - | |
| DETR[28] | 96 | 93 | 96 | 94 | |
| Proposed | 99.78 | 99.78 | 99.78 | 99.78 | |
| IQ-QTH/NCCD Dataset | InceptionNeXT_ViT[19] | 99.54 | 99.67 | 99.6 | 99.12 |
| CNN+DA [8] | 98.78 | 97.57 | 97.67 | 98.78 | |
| DenseNet [8] | 94.1 | 93.8 | 94 | 94.2 | |
| ResNet [8] | 93 | 92.5 | 93.1 | 92.7 | |
| EfficientNet B0 [8] | 92.64 | 91.81 | 91.78 | 91.21 | |
| FocalNeXT [20] | 99.81 | 99.78 | 99.36 | 99.56 | |
| ResNet 50 [26] | 93.15 | 88.51 | 86.29 | 87.29 | |
| SwinBase [26] | 97.72 | 94.25 | 97.99 | 95.82 | |
| Max ViT-Base [19] | 97.26 | 96.5 | 93.75 | 94.97 | |
| DieT3 Base [19] | 98.17 | 96.42 | 96.42 | 96.42 | |
| Proposed | 99.94 | 99.96 | 99.96 | 99.96 |
Traditional CNN-based models such as VGG-16, ResNet-50, Custom CNN, VER-Net, ResNet101+Inception V3, and hybrid CNN architectures demonstrated moderate to strong performance, with accuracies ranging from 78.80% to 95.80%. More advanced architectures like Xception (95.80%) and VGG19+LSTM (99.42%) improved accuracy but could not simultaneously achieve balanced precision, recall, and F1-scores. Transformer-based approaches such as Vision Transformer (94%) and DETR (96%) provided better generalization, yet still fell short of optimal overall performance. In contrast, the proposed HierLungXAI framework achieved 99.78% across all metrics accuracy, precision, recall, and F1-score demonstrating superior performance by effectively capturing both fine-grained nodule details and broader contextual dependencies through hierarchical local–global feature extraction.
Similarly, on the IQ-QTH/NCCD dataset, competing models such as DenseNet (94.10%), ResNet (93.00%), EfficientNet-B0 (92.64%), and ResNet-50 (93.15%) yielded lower performance, while advanced vision transformers like SwinBase (97.72%), MaxViT-Base (97.26%), and DeiT3-Base (98.17%) showed stronger results but still fell short of optimal consistency. Even recent high-performing methods including InceptionNeXT_ViT (99.54%) and FocalNeXT (99.81%) could not surpass the proposed framework, which achieved 99.94% accuracy and 99.96% precision, recall, and F1-score. This superior performance can be attributed to the AISEP preprocessing pipeline, which ensures standardized and noise-reduced CT slices, combined with the HierEffNet architecture that integrates local feature learning with hierarchical attention for global context modeling. Additionally, by concentrating on pertinent diseased regions, Grad- CAM-based interpretability guarantees that forecasts are not only accurate but also clinically useful. Overall, the findings show how innovative and reliable the suggested framework is, outperforming both transformer-based and CNN-based designs on a variety of datasets.
5.4 Ablation Study
An ablation research was conducted using the Chest CT Image Dataset and the IQ-QTH/NCCD Dataset in order to assess the contribution of each element in the suggested HierLungXAI framework. The evaluation followed a progressive structure, beginning with a Baseline CNN trained on raw CT images to establish a reference point. Introducing the AISEP (Advanced Image Standardization and Enhancement Pipeline) improved accuracy and precision significantly, stressing the significance of noise reduction, contrast improvement, and systematic intensity normalization for constant image quality. The addition of HierEffNet made it possible to extract fine-grained local characteristics, capturing subtle nodular details that were previously overlooked, which translated into noticeable gains across all metrics. The incorporation of the Hierarchical Attention Module (HAM) further enhanced the framework by modeling volumetric context, inter-slice dependencies, and global structural patterns, leading to significant improvements in recall and F1-score. Finally, the complete integration of all components with Grad-CAM explainability not only delivered the best performance across accuracy, precision, recall, and F1-score but also ensured transparency by visually highlighting the critical regions influencing malignancy predictions.
As summarized in Table 10, each component contributes distinctively to performance improvements.
| Configuration | Dataset | Accuracy (%) | Precision (%) | Recall (%) | F1-score (%) |
| Baseline CNN (without AISEP) | Chest CT | 92.1 | 91.85 | 92.4 | 92.12 |
| CNN+ AISEP | 95.84 | 95.62 | 95.9 | 95.71 | |
| AISEP + HierEffNet | 97.62 | 97.48 | 97.66 | 97.56 | |
| AISEP + HierEffNet + HAM | 99.02 | 99 | 99.04 | 99.02 | |
| Proposed (+ AISEP + HierEffNet + HAM + Grad-CAM) | 99.78 | 99.78 | 99.78 | 99.78 | |
| Baseline CNN (without AISEP) | IQ-QTH/NCCD | 93.25 | 92.84 | 93.3 | 93.07 |
| CNN+ AISEP | 96.72 | 96.41 | 96.8 | 96.6 | |
| AISEP + HierEffNet | 98.34 | 98.4 | 98.3 | 98.35 | |
| AISEP + HierEffNet + HAM | 99.42 | 99.46 | 99.4 | 99.43 | |
| Proposed (+ AISEP + HierEffNet + HAM + Grad-CAM) | 99.94 | 99.96 | 99.96 | 99.96 |
The baseline CNN demonstrated moderate capability but remained sensitive to imaging variability. AISEP contributed to data consistency, HierEffNet strengthened sensitivity to small nodules, HAM captured inter-slice contextual information, and Grad-CAM introduced interpretability, together forming a comprehensive and clinically reliable pipeline. These results confirm that the synergy between preprocessing, hierarchical multi-level feature extraction, contextual attention modeling, and explainability drives the superior accuracy and robustness of the proposed HierLungXAI framework, achieving nearly perfect performance across both datasets and setting a new benchmark for CT-based LC classification.
5.5. Discussion
The proposed HierLungXAI framework was designed to overcome the persistent challenges of CT-based LC classification, where traditional methods often suffer from inconsistent preprocessing, noise interference, subtle nodule visibility, and inadequate modeling of inter-slice dependencies. A key novelty of this work lies in unifying preprocessing, hierarchical multi-scale feature extraction, volumetric attention modeling, and explainability into a single clinically driven pipeline. Unlike prior CNN- or transformer-based approaches that emphasize either localized or global features in isolation, HierLungXAI ensures balanced representation of both fine-grained nodular details and long-range volumetric context. The AISEP preprocessing pipeline represents a major innovation, as it standardizes intensity distributions, enhances contrast, reduces noise, and ensures spatial consistency, directly addressing the variability in CT acquisition protocols. By improving image quality and preserving subtle nodular characteristics, AISEP provides a robust foundation for reliable feature extraction. The HierEffNet module introduces a hierarchical design that captures nodule-specific details while maintaining computational efficiency, thereby enhancing sensitivity to small and low-contrast nodules. Complementing this, the Hierarchical Attention Module (HAM) incorporates inter-slice dependencies and volumetric relationships, enabling the framework to learn broader global context and structural variations critical for accurate classification. The contribution of each component was validated through an ablation study. Baseline CNN models showed moderate performance and sensitivity to imaging variability. With AISEP, significant gains in accuracy and precision were observed, confirming the importance of robust preprocessing. Adding HierEffNet further boosted feature learning, particularly for subtle nodules, while HAM improved recall and F1-score by capturing global dependencies. Finally, the complete integration with Grad-CAM not only achieved the best performance across all metrics (99.78% on Chest CT and 99.94% on IQ-QTH/NCCD) but also provided transparent visual explanations, enhancing clinical trust. Overall, the novelty of HierLungXAI lies in its systematic integration of image standardization, hierarchical feature extraction, global attention, and interpretability into a single pipeline. In addition to achieving cutting-edge accuracy, this comprehensive design closes the gap between algorithmic forecasts and clinically reliable decision-support, underscoring its significance for real-world adoption.
5.6. Limitations and Generalizability
Although the proposed HierLungXAI framework demonstrates superior performance on the Chest CT and IQ-OTH/NCCD benchmark datasets, certain limitations should be acknowledged. First, despite being publicly available and widely used in prior studies, the employed datasets are relatively limited in size and may not fully capture the heterogeneity encountered in real-world clinical environments. In particular, variations arising from multi-institutional data sources, different CT scanner vendors, acquisition protocols, slice thicknesses, and reconstruction settings are not comprehensively represented in these datasets.
Second, the datasets primarily consist of well- curated and pre-labeled CT images, which may differ from routine clinical scans that often contain motion artifacts, incomplete lung coverage, or ambiguous nodule boundaries. While the proposed AISEP preprocessing pipeline improves robustness against noise and intensity variability, the generalizability of the framework to large-scale, multi-center clinical cohorts requires further validation.Finally, although the use of hierarchical feature extraction and attention-based volumetric modeling enhances sensitivity to subtle nodules, the current evaluation is restricted to retrospective datasets. Prospective validation and external testing on independent multi-institutional cohorts are necessary to fully establish clinical reliability.Future work will focus on extending HierLungXAI to larger and more diverse datasets, including multi-center and multi-scanner CT collections, and exploring domain adaptation strategies to further improve generalization. Additionally, real-time clinical validation and integration into radiology workflows will be investigated to assess practical deployment feasibility. In conclusion, this research presented HierLungXAI, a novel and comprehensive framework for accurate and interpretable LC detection from CT scans. Unlike existing approaches that often emphasize either localized features or global context in isolation, the proposed system integrates a standardized preprocessing pipeline, hierarchical feature extraction, volumetric attention modeling, and explainable AI into a unified workflow. The novelty of this framework lies in its holistic design: the AISEP module ensures consistent input quality by addressing intensity variability, noise, and spatial inconsistencies; HierEffNet captures fine-grained nodule-specific details while maintaining computational efficiency; and the Hierarchical Attention Module effectively models inter-slice dependencies, ensuring robust representation of volumetric structures. By coupling these with Grad-CAM interpretability, the framework not only achieves state-of-the-art accuracy but also provides transparent decision support, fostering greater clinical trust. Experimental results demonstrated the superiority of HierLungXAI across two benchmark datasets, achieving 99.78% accuracy on the Chest CT dataset and 99.94% accuracy on the IQ-QTH/NCCD dataset, with near-perfect scores across all evaluation metrics. The ablation study further validated the incremental impact of each component, confirming the necessity of integrating preprocessing, hierarchical learning, attention modeling, and interpretability for optimal performance. While the current framework demonstrates exceptional accuracy and robustness, its reliance on hierarchical CNN-based modules may limit adaptability to highly diverse and multi-scale nodules across heterogeneous datasets. Future enhancements will explore hybrid CNN Transformer architectures that combine efficient local feature extraction with transformer-based global attention, enabling richer contextual modeling and improved generalization. Such advancements are expected to further enhance robustness, interpretability, and clinical readiness of automated LC detection systems.
Declarations
Ethics approval and consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
All contributors agreed and given consent to Publication.
Availability of data and material
The datasets used in this study are publicly available, including the Chest CT dataset from Kaggle and the IQ- OTH/NCCD dataset. No private or confidential patient data were used.
Competing interests
On behalf of all authors, the corresponding author states that they have no competing interest.
Funding
No fund was received for this work.
Authors contributions
All authors are equally contributed.
References
- Recent advances in lung cancer research: unravelling the future of treatment Bertolaccini L, Casiraghi M, Uslenghi C, Maiorca S, Spaggiari L. Updates in Surgery.2024;76(6). CrossRef
- Radiomics as a non-invasive adjunct to Chest CT in distinguishing benign and malignant lung nodules Selvam M, Chandrasekharan A, Sadanandan A, Anand VK , Murali A, Krishnamurthi G. Scientific Reports.2023;13(1). CrossRef
- A methodical exploration of imaging modalities from dataset to detection through machine learning paradigms in prominent lung disease diagnosis: a review Kumar S, Kumar H, Kumar G, Singh SP , Bijalwan A, Diwakar M. BMC medical imaging.2024;24(1). CrossRef
- Eliminating the stigma: A systematic review of the health effects of low-dose radiation within the diagnostic imaging department and its implications for the future of medical radiation Mack SA . Journal of Medical Imaging and Radiation Sciences.2020;51(4). CrossRef
- Lung Nodule Detection from Feature Engineering to Deep Learning in Thoracic CT Images: a Comprehensive Review Halder A, Dey D, Sadhu AK . Journal of Digital Imaging.2020;33(3). CrossRef
- Interpretable classification of computed tomography scans for identification of lung cancer BRUSCHI G. 2021.
- Deep Learning for Medical Image-Based Cancer Diagnosis Jiang X, Hu Z, Wang S, Zhang Y. Cancers.2023;15(14). CrossRef
- Optimizing non small cell lung cancer detection with convolutional neural networks and differential augmentation Shariff V, Paritala C, Ankala KM . Scientific Reports.2025;15(1). CrossRef
- Explainable Ai For Healthcare: Developing Transparent And Interpretable Models For Medical Diagnosis Kenigson J. 2025.
- Artificial Intelligence in Detection and Classification of Lung Cancer ‐ An Overview Chakraborty S, Banerjee DK . .
- Deep Learning Techniques for Lung Cancer Diagnosis with Computed Tomography Imaging: A Systematic Review for Detection, Segmentation, and Classification Abdullahi K, Ramakrishnan K, Ali AB . Information.2025;16(6). CrossRef
- A Review of Deep Learning Techniques for Lung Cancer Screening and Diagnosis Based on CT Images Thanoon MA , Zulkifley MA , Mohd Zainuri MAA , Abdani SR . Diagnostics.2023;13(16). CrossRef
- Explainability, transparency and black box challenges of AI in radiology: impact on patient care in cardiovascular radiology Marey A, Arjmand P, Alerab ADS , Eslami MJ , Saad AM , Sanchez N, Umair M. Egyptian Journal of Radiology and Nuclear Medicine.2024;55(1). CrossRef
- Explainable AI for lung cancer detection via a custom CNN on CT images Hammad M, ElAffendi M, El-Latif AAA , Ateya AA , Ali G, Plawiak P. Scientific Reports.2025;15(1). CrossRef
- Advanced U-Net Architectures with CNN Backbones for Automated Lung Cancer Detection and Segmentation in Chest CT Images Golkarieh A, Kiashemshaki K, Boroujeni SR , Isakan NA . 2025. CrossRef
- AI-Powered Lung Cancer Detection: Assessing VGG16 and CNN Architectures for CT Scan Image Classification Klangbunrueang R, Pookduang P, Chansanam W, Lunrasri T. Informatics.2025;12(1). CrossRef
- VER-Net: a hybrid transfer learning model for lung cancer detection using CT scan images Saha A, Ganie SM , Pramanik PKD , Yadav RK , Mallik S, Zhao Z. BMC Medical Imaging.2024;24(1). CrossRef
- Lung cancer detection and severity analysis with a 3D deep learning CNN Model using CT-DICOM clinical dataset Eldho KJ , Nithyanandh S. Indian J Sci Technol.2024;17(10):899-910. CrossRef
- Attention Enhanced InceptionNeXt-Based Hybrid Deep Learning Model for Lung Cancer Detection Ozdemir B, Aslan E, Pacal I. IEEE Access.2025;13. CrossRef
- FocalNeXt: A ConvNeXt augmented FocalNet architecture for lung cancer classification from CT-scan images Gulsoy T, Baykal Kablan E. Expert Systems with Applications.2025;261. CrossRef
- Lung cancer detection via deep learning-based pyramid network with honey badger algorithm Angel mary A., Thanammal K. K.. Measurement: Sensors.2024;31. CrossRef
- Stacked neural nets for increased accuracy on classification on lung cancer Br SRR , Sen S, Bhatt R, Dhanetwal ML , Sharma M, Naaz R. Measurement: Sensors.2024;32. CrossRef
- A CAD System for Lung Cancer Detection Using Hybrid Deep Learning Techniques Alsheikhy AA , Said Y, Shawly T, Alzahrani AK , Lahza H. Diagnostics.2023;13(6). CrossRef
- Lung cancer detection using ensemble technique of CNN. In Proceedings of International Conference on Information and Communication Technology for Development: ICICTD 2022 (pp. 497-507). Singapore: Springer Nature Singapore Akhand ZEN , Rahman AI , Sarda A, Fahim MZA , Tushi LT , Azad K, Tahiat HT . 2023.
- Lcdctcnn: Lung cancer diagnosis of ct scan images using cnn based model. In 2023 10th International Conference on Signal Processing and Integrated Networks (SPIN) (pp. 205-212) Mamun M, Mahmud MI , Meherin M, Abdelgawad A. IEEE.2023.
- Classification of non-small cell lung cancers using deep convolutional neural networks Atiya SU , Ramesh N. V. K., Reddy BNK . Multimedia Tools and Applications.2024;83(5). CrossRef
- A novel approach to integrating Vision Transformers and machine learning for robust lung nodule classification using CT imaging Huang Y, Li Q, Chen H, Zhou X, Han Q, Lu H, Cui N, Cao Q. Journal of Radiation Research and Applied Sciences.2025;18(3). CrossRef
- A transformer‐based deep neural network for detection and classification of lung cancer via PET/CT images Barbouchi K, El Hamdi D, Elouedi I, Aicha TB , Echi AK , Slim . ResearchGate.2023. CrossRef
License

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Copyright
© Asian Pacific Journal of Cancer Biology , 2026
Author Details