Molecular Foundations of Cancer: A Narrative Review of Hallmarks, Microenvironment, and Precision Therapeutic Advances
Download
Abstract
Background: Cancer represents a devastating global health challenge characterized by intricate molecular mechanisms that defy conventional therapeutic approaches. This review integrates contemporary scientific understanding of the biological processes underlying cancer, exploring the complex interplay among genetic, cellular, and microenvironmental factors that drive tumor development. It critically examines pivotal hallmarks such as dysregulated cellular proliferation, metastatic potential, and immune system interactions. Central molecular signaling pathways, including MAPK and PI3K-AKT, are analyzed to reveal the dynamics of malignant transformation. Furthermore, the exploration of the tumor microenvironment and cellular metabolic adaptations illustrates the resilience of cancer cells. The aim of this narrative review is to synthesize foundational molecular mechanisms and emerging precision therapeutic strategies to provide a comprehensive perspective on the molecular complexity of cancer.
Conclusion: Emerging therapeutic strategies represent a paradigm shift towards precision medicine, leveraging targeted molecular interventions, immunotherapy, and advanced nanotechnology. This review provides a unique synthesis by specifically linking foundational mechanisms, such as epigenetic modifications, to the pathological heterogeneity observed in tumors. By integrating current scientific knowledge and technological innovations like CAR-T therapies, this work provides a comprehensive perspective on the future of personalized cancer management.
1. Introduction
Cancer is a multifaceted disease that causes over 10 million deaths annually, worldwide. It is characterised by diverse biological traits and a wide range of phenotypes [1, 2]. Breast cancer, the most prevalent malignant tumor among women, exemplifies this heterogeneity through its varied cellular states and interactions [3]. This complexity arises from the interplay of genetic, environmental, and host factors, resulting in biological heterogeneity that influences the pathophysiology, tumor biology, and clinical outcomes [4]. Research indicates that bulk tumors often originate from “cancer stem cells” (CSCs), which are responsible for cancer initiation and recurrence [1]. Epigenetic mechanisms are integral to cancer development, alter cellular states, and play a significant role in tumour origin and progression [1].
The global impact and complexity of cancer necessitate ongoing molecular research to enhance clinical care and develop personalised treatments. Advanced technologies, such as single-cell RNA sequencing (scRNA-seq), are revolutionising cancer research by providing insights into cellular heterogeneity [3]. Understanding molecular characteristics at the genomic level is crucial for improving clinical care and developing personalised precision medicine approaches for patients with cancer [4]. The aim of this narrative review is to synthesize the foundational molecular mechanisms, the complexities of the tumor microenvironment, and emerging precision therapeutic strategies to provide a comprehensive perspective on the current landscape of oncology.
2. Cellular Foundations of Cancer
Normal cell cycle and regulation
The eukaryotic cell cycle involves the synthesis and degradation of cyclins that activate cyclin-dependent kinases (CDKs) [5]. Key proteins, including cyclins, CDKs, and the inhibitors p21 and p16, control cell proliferation at checkpoints [5]. These checkpoints function as DNA surveillance mechanisms during cell division [6]. In response to damage, checkpoints can delay progression or induce cell cycle exit or apoptosis [6]. CDC25 phosphatases are crucial in cell cycle phase transitions and are targets of the checkpoint machinery [7]. Meiotic cycles have checkpoints that ensure recombination completion before spindle formation and proper chromosomal attachment. Animal oocytes maintain meiotic arrest via cytostatic factors [8]. Cell cycle regulation occurs through temporal and spatial information, including the synthesis and activation of proteins that regulate gene expression. Checkpoints link cell division to chromosome states, whereas regulatory components direct organelle positioning [9]. Understanding these mechanisms is crucial for developing cancer therapies and addressing cell cycle disorders.
Fundamental cellular alterations in cancer
Cancer development involves cellular modifications, including genetic disruption and molecular changes, that enable malignant progression. Genetic inactivation of tumor suppressor genes such as p16 (INK4a)/p14 (ARF) and NF2 and epigenetic silencing of genes such as RASSF1A are prevalent in cancers. The activation of receptor tyrosine kinases and deregulation of the MAPK and PI3K-AKT signalling pathways contribute to cancer cell survival [10]. Although specific oncogene mutations, such as KRAS, are common in certain cancers, they are not universal [10, 11]. Cancer is characterised by dysregulation of pathways that drive aberrant cell growth. Cellular transformation involves oncogene activation and tumour suppressor inactivation [11]. The interaction between genetic mutations, epigenetic changes, and altered signalling contributes to cancer progression. Understanding these processes is vital for developing targeted therapies and improving treatment.
Hallmarks of cancer development
Cancer progression is characterized by uncontrolled growth and systemic transformation. The framework of Hanahan and Weinberg (2000,) [12] explains the complexity and adaptability of cancer [13, 14]. Core hallmarks include sustained proliferative signalling, evasion of growth suppressors, resistance to cell death, replicative immortality, angiogenesis, and metastasis [13, 15, 16]. Emerging hallmarks include dysregulated cellular energetics, immune evasion, genome instability, and tumor-promoting inflammation [14]. The AKT pathway influences multiple hallmarks, thereby affecting cellular physiology and oncogenesis [14]. TGF-β functions as a tumor suppressor in normal cells and promotes tumor growth in malignant cells [16]. Understanding these hallmarks is crucial for developing targeted therapies. The multifaceted nature of cancer requires comprehensive approaches addressing tumor biology [17, 18] (Table 1).
| Hallmark | Key Molecular Mechanism(s) | Example Therapeutic Class / Drug |
| Sustained Proliferative Signalling | Oncogene activation (e.g., EGFR, HER2, BRAF) | Tyrosine Kinase Inhibitors (e.g., Osimertinib, Trastuzumab, Dabrafenib) |
| Evasion of Growth Suppressors | Inactivation of RB pathway | CDK4/6 Inhibitors (e.g., Palbociclib) |
| Resistance to Cell Death | Overexpression of anti-apoptotic proteins (e.g., BCL-2) | BH3 Mimetics (e.g., Venetoclax) |
| Replicative Immortality | Telomerase activation | Telomerase Inhibitors (e.g., Imetelstat) |
| Induction of Angiogenesis | Upregulation of VEGF | |
| Activation of Invasion & Metastasis | EMT activation; HGF/c-MET signalling | MET Inhibitors (e.g., Crizotinib) |
| Deregulating Cellular Energetics | Warburg effect; IDH mutations | IDH Inhibitors (e.g., Ivosidenib); Glycolysis inhibitors (experimental) |
| Genome Instability & Mutation | Defective DNA Damage Response (DDR) (e.g., BRCA mutations) | PARP Inhibitors (e.g., Olaparib) |
| Avoiding Immune Destruction | Expression of immune checkpoints (e.g., PD-L1) | Immune Checkpoint Inhibitors (e.g., Pembrolizumab, Nivolumab) |
| Tumor-Promoting Inflammation | Inflammatory signalling (e.g., NF-κB) | Anti-inflammatory drugs (e.g., IL-6 inhibitors, JAK inhibitors) |
3. Genetic and Molecular Basis of Cancer
Genomic instability
Genomic instability, a defining characteristic of cancer, arises from chromosomal aberrations, failures of the DNA damage response (DDR), and structural alterations. DNA damage can result from endogenous sources, such as replication errors, or exogenous factors, such as radiation and chemicals, leading to nucleotide changes or chromosomal abnormalities [19]. Unrepaired damage disrupts cellular homeostasis, activates oncogenes, inactivates tumor suppressors, and drives carcinogenesis [19, 20]. Chromosomal instability encompasses karyotypic changes, copy-number alterations, and structural abnormalities [20]. DNA-damaging events, such as chromothripsis and chromoplexy, can cause extensive genomic insults and exacerbate genomic instability in cancers [21]. Defects in the DDR pathway can lead to aberrant DNA repair and tumorigenesis [22]. Dysfunctional telomeres trigger DDR activation, causing chromosome-end fusion and genomic instability [22]. Abnormalities in DDR-associated genes and dysfunctional ubiquitin-proteasome systems are prevalent in metastatic castration-resistant prostate cancer [23]. Understanding these mechanisms is crucial for developing novel prognostic biomarkers and targeted therapies [21, 23]. Clinically, instability can be assessed by using histopathological tools. Immunohistochemistry (IHC) for mismatch repair (MMR) proteins is a standard method for screening Lynch syndrome and has become a critical pan-tumor biomarker for immune checkpoint inhibitor responses [24]. Loss of nuclear staining for these proteins provides a visual correlation to genomic defects and has therapeutic implications [25].
Oncogenes and tumor suppressor genes
Oncogene activation and tumor suppressor gene silencing are pivotal for cancer initiation and progression. Oncogenes, which are proto-oncogenes involved in cell growth, can be activated by gene amplification, chromosomal rearrangements, or mutations [26, 27]. This activation led to uncontrolled cell proliferation. Tumor suppressor genes that inhibit cell transformation and regulate cell cycle control and apoptosis are inactivated in cancer cells through mutations, deletions, or epigenetic silencing [28, 29]. Studies have shown a correlation between oncogene activation and epigenetic silencing of specific tumor suppressor genes [30]. The transcriptional repressor Id1, which is overexpressed in numerous cancers, targets tumor suppressor genes such as TSP1 and CDKN2A (p16). The precise mechanism of this silencing remains unclear, as Id1 overexpression alone does not repress these target genes in melanoma cell lines [30] (Healey et al., 2010). This suggests that oncogene activation may require additional genetic or epigenetic alterations to silence the tumor suppressor genes. The interplay between oncogene activation and silencing of tumor suppressor genes is complex. Understanding these mechanisms is crucial to developing novel cancer therapies. Emerging techniques, such as RNA interference, zinc finger nucleases, and CRISPR, offer promising avenues for targeting these molecular pathways in cancer treatment [26].
Genetic mutations and cancer progression
Driver mutations confer a selective advantage to cancer cells, while passenger mutations are considered neutral or mildly harmful [31]. Drivers are the primary contributors to carcinogenesis, while passengers have minimal impact. Recent studies have shown complex interactions among these mutations during cancer evolution. Computational models suggest that passenger mutations can accelerate cancer evolution by reducing population fitness and boosting the fitness of intermediate mutants in tumor suppressor gene inactivation pathways [32, 33]. This suggests that the baseline rate of tumor suppressor gene inactivation may be faster than previously thought. Although some passenger mutations harm cancer cells, their cumulative effect creates a barrier to cancer progression, defined by critical population-size and mutation-rate thresholds [34]. The genetic evolution of cancer involves balancing the driver and passenger mutations. Although drivers mainly propel cancer progression, passenger accumulation can significantly affect tumor fitness and treatment outcomes. This understanding has implications for cancer therapy, suggesting that targeting the collective burden of passenger mutations is promising [34, 35]. The timing and dependencies of mutations, including low-frequency drivers, are crucial in cancer genomics [36].
Epigenetic modifications
Epigenetic modifications include DNA methylation, histone modifications, chromatin remodeling, gene regulation, and cancer development. DNA methylation, particularly promoter hypermethylation, can silence tumor suppressor genes and contribute to cancer progression [37]. Histone modifications affect chromatin structure and gene expression, with aberrant patterns found in various tumor types [38, 39]. Interactions between epigenetic mechanisms increase the complexity of gene regulation in cancer. Crosstalk between N6-methyladenosine (m6A) RNA methylation and other epigenetic modifications initiates epigenetic remodeling and influences tumorigenesis [40]. The relationship between histone modifications and regulatory elements in breast cancer cells shows that active regulatory elements are associated with active histone modifications near transcription start sites [41]. Dysregulation of epigenetic mechanisms contributes to cancer development. The reversible nature of these modifications has led to epigenetic therapies targeting the enzymes involved in DNA methylation and histone modifications [37, 38]. Understanding the interplay between epigenetic mechanisms and their roles in gene regulation is crucial for developing effective cancer treatments and advancing the knowledge of tumor biology [42].
Genomic landscape of cancer
Genomic analyses of different cancer types have revealed a landscape of genetic alterations, including mutations, copy number changes, and structural variations. Whole-genome sequencing of 403 adult gliomas has identified recurrent coding and non-coding mutations, providing a map of structural variants and copy-number changes [43]. Analysis of 33,036 solid tumors showed that copy number changes in cancer driver genes are broadly prognostic, whereas mutations are rarely associated with patient survival [44]. There are discrepancies in the prognostic value of genetic alterations. Although mutations in cancer driver genes do not correlate with patient outcomes, copy number changes and methylation patterns are more informative for risk assessment [44]. The genetic landscape varies among the cancer types. RHOA mutations were observed in 14.3% of diffuse-type gastric tumors but not in intestinal-type tumors [45]. The genomic terrain of cancer shows heterogeneity across cancer types. Mutations, copy-number alterations, and structural variations contribute to molecular signatures. Integrating genomic, epigenomic, and transcriptomic profiles provides an understanding of cancer biology and may improve diagnostic, prognostic, and therapeutic strategies [45, 46]. This approach offers insight into personalized medicine and targeted therapies. In clinical practice, Next-Generation Sequencing (NGS) panels are used to analyze tumor genomics. The data, identifying driver mutations, copy number alterations, and fusions, are discussed by molecular tumor boards (MTBs) to recommend targeted therapies or determine prognosis. Integrating genomic data into clinical decision-making is essential for precision oncology [47].
4. Molecular Signalling in Cancer Development
Cell proliferation signalling pathways
Protein kinases play a pivotal role in cellular signalling, governing cell growth, proliferation, and survival [48]. These networks encompassing numerous kinases and phosphorylation sites are frequently dysregulated in cancer. MAPK cascades convey signals to intracellular targets and initiate cell proliferation, differentiation, and apoptosis [49]. In cancer, mutations or aberrant protein expression can disrupt these networks uncontrolled cellular growth. Ras G-proteins, which regulate MAPK pathways, are mutated in approximately 30% of human cancers [50], highlighting the need for targeted therapies. Research shows more kinases than previously recognized contribute to tumorigenesis, suggesting new therapeutic strategies [48]. The complexity of these networks is demonstrated by pathway interactions; cAMP can modulate growth factor responses by decoupling MAPK cascades and activating signals via PKA [51]. Understanding these interactions and molecular alterations in signalling pathways is essential for elucidating cancer growth circuits and developing effective therapies.
Apoptosis and cell survival mechanisms
Cancer cells employ molecular strategies to circumvent cell death, a fundamental characteristic of malignancy, by suppressing apoptosis and resistance to pro-death signals [52]. Dysregulated apoptosis contributes to cancer progression and resistance to treatments such as radiation and cytotoxic drugs [53]. Cancer cell survival depends on the overexpression of anti-apoptotic proteins, such as Bcl-2, which confer resistance to apoptosis-inducing agents and are activated in lymphomas and other cancers [53]. Resistance develops through mutations in pro-apoptotic genes, increased expression of anti-apoptotic proteins, and the silencing of pro-apoptotic genes via DNA hypermethylation [54]. Studies have shown that endocytosis plays a key role in apoptotic resistance and immune escape, with tumors exhibiting dysregulated endocytic dynamics that support metabolic demands and resistance to therapy [55]. Alternative cell death pathways, such as ferroptosis, entosis, necroptosis, and immunogenic cell death, present new intervention opportunities [56]. Cancer cells resist death by disrupting apoptotic pathways and altering endocytic dynamics. Understanding these mechanisms is crucial for developing therapies targeting the apoptotic machinery and alternative cell death mechanisms for effective cancer treatment [55-57].
Growth factor signalling
Receptor tyrosine kinases (RTKs) are integral to cellular communication through growth factor signalling. These transmembrane proteins are activated upon ligand binding, inducing receptor dimerization and autophosphorylation of tyrosine residues within the intracellular domains [58]. This activation triggers signal transduction cascades involving adaptor proteins and enzymes that regulate cellular processes, including growth, differentiation, metabolism, and motility [59]. RTKs can also translocate to the nucleus to initiate transcription [60]. Proteomic studies have identified novel post-translational modification sites, highlighting the complexity of these networks [61]. Aberrant RTK signalling occurs during cancer progression through receptor overexpression, mutations, gene amplification, and defective regulation [60, 62]. These changes activate pathways, such as MAPK, PI3K/Akt, and JAK/STAT, promoting cancer stemness, angiogenesis, and metastasis [62]. Although RTKs are attractive therapeutic targets, resistance and toxicity challenge RTK-targeted therapies [63], underscoring the need for new approaches to modulate these pathways in cancer treatment.
Cellular communication and transformation
Cancer cells interact through molecular transformations and signalling networks to establish tumor progression and immune evasion [64]. These interactions occur via extracellular vesicle (EVs)-transferring of proteins, nucleic acids, and microRNAs between cells [65]. The tumor microenvironment (TME) enables intercellular communication, in which tumor-associated macrophages (TAMs) maintain immunosuppression through EVs [66]. Adipocytes in the breast stroma secrete EVs that promote tumor growth [65]. Tunneling nanotubes (TNTs) enable direct intercellular transport, which affects cellular reprogramming and chemotherapy resistance [67]. Cancer cells use these pathways to modify their TME. Understanding these interactions is vital to developing novel therapeutic strategies. Single-cell RNA sequencing and tools such as CellChat have enhanced our understanding, potentially leading to improved anticancer therapies [68-71].
Immune system evasion mechanisms
Cancer cells utilize molecular mechanisms to evade immune surveillance and resist immune checkpoint inhibitors (ICIs). These mechanisms include the modulation of immune checkpoints, molecular alterations, and systemic immune resistance. Cancer cells evade immune attacks by modulating immune checkpoints, with tumors upregulating inhibitory checkpoints, such as PD-1/ PD-L1 and CTLA-4, to suppress T cell activity [72, 73].
Alternative immune checkpoint activation contributes to resistance to PD-1/PD-L1 inhibitors [74]. The triggering receptor TREM2 negatively affects antitumor immunity by inhibiting T cell proliferation [75]. Cancer cells evade detection through neoantigen loss or impaired antigen presentation [74]. Epithelial-mesenchymal transition facilitates immune evasion by altering the expression of checkpoint molecules [76]. Epigenetic modifications alter the expression of immune response genes and contribute to drug resistance [74]. Cancer cells create an immunosuppressive tumor microenvironment by reprogramming their metabolism to generate nutrient scarcity and hypoxia [77] (Jiang et al., 2020). “Cold” tumors lacking T cell infiltration render ICI therapy ineffective [78], while gut microbiome dysbiosis contributes to resistance [74]. Understanding these mechanisms is crucial to developing strategies to overcome resistance. Solutions include combining ICIs with alternative checkpoint therapies and enhancing T cell infiltration [72, 78].
5. Cancer Microenvironment
Tumor microenvironment composition
The cellular landscape of cancer is defined by interactions between cancer cells and tumor microenvironment (TME) components, forming a dynamic ecosystem that supports cancer cell survival and growth [79, 80]. Cancer cells interact with stromal, endothelial, and immune cells, creating an ecosystem that facilitates tumor progression and treatment resistance [80]. TME comprises immune cells, stromal cells, and soluble factors that interact with cancer cells [79]. This crosstalk shapes the immune landscape and influences tumor progression and therapeutic response [79, 81]. Cancer-associated fibroblasts (CAFs) undergo phenotypic changes that affect tumor growth, promote cancer cell survival, and facilitate immune evasion [82]. The extracellular matrix (ECM) regulates cell growth, survival, and migration through cellular interactions [83]. Cancer cell plasticity enables environmental adaptation, contributing to tumor diversity and drug resistance through processes such as epithelial-mesenchymal transition [84]. Metabolic interplay within the TME supports cancer cell proliferation [82]. Understanding the interactions and mechanisms underlying tumor heterogeneity and drug resistance is crucial for developing effective cancer therapies [79, 81, 84].
Role of stromal cells
The tumor microenvironment (TME) is integral to cancer progression and metastasis through interactions between cancer cells, stromal cells, and extracellular matrix (ECM) components. These interactions form communication networks that support tumor growth and invasion [85-87]. Cancer-associated fibroblasts (CAFs) are crucial within the TME for synthesizing and modulating the ECM, which serves as a scaffold for growth factors and cytokines that promote tumor growth and metastasis [88]. ECM components regulate growth factors, such as hepatocyte growth factor (HGF) and its receptor c-MET, influencing cellular processes, including proliferation and angiogenesis [86]. Gap junctions (GJs) show context- dependent effects on cancer progression, potentially inhibiting metastasis in some models, while promoting epithelial-to-mesenchymal transition (EMT) in breast cancer [89]. Stromal dynamics of cancer progression involve complex communication networks and molecular pathways. Understanding these interactions is crucial for developing TME-targeted therapies and for improving cancer management [90, 91].
Inflammation and cancer progression
Inflammation plays a dual role in cancer, functioning as both a protective mechanism and a potential catalyst for tumorigenesis. Acute inflammation defends against pathogens and tissue damage, whereas chronic inflammation can facilitate malignant transformation [92]. The inflammatory microenvironment is regulated by mediators, including cytokines (TNF-α, IL-6, TGF-β, and IL-10), chemokines, and growth factors, which contribute to tumor progression [93]. A bidirectional relationship exists between inflammation and cancer; chronic inflammation can lead to cancer, and cancer cells can induce inflammatory responses that support tumor development. Breast cancer cells produce TSLP, triggering dendritic cells to express OX40L, which in turn drives the development of inflammatory Th2 cells that promote tumor growth [94]. The IL-6 amplifying mechanism, involving NF-κB and STAT3, sustains chronic inflammation and drives tumorigenesis [95]. Molecular triggers such as PAMPs and DAMPs initiate chronic inflammatory responses [96], creating conditions for tumor development by inducing DNA damage, promoting cell proliferation, and suppressing antitumor immunity. Understanding these interactions is essential for developing targeted therapies against inflammation- associated malignancies.
Angiogenesis
Blood vessel formation involves multiple molecular switches that are critical for tumor nutrient acquisition and growth, including vasculogenesis and angiogenesis [97, 98]. Hypoxia and the hypoxia-inducible factor (HIF) transcriptional system trigger blood vessel growth in malignant tumors [99]. HIF-1α stabilization under hypoxic conditions increases the transcription of proangiogenic genes, particularly vascular endothelial growth factor (VEGF) [98]. The angiogenic switch involves transitioning from a pre-vascular to a vascularized tumor phenotype, which is controlled by pro- and anti-angiogenic factors [100]. Vascular remodeling integrates existing vessels into tumor vasculature and vessel regression, transforming healthy networks into tumor-specific vasculature [101]. This process is influenced by molecular pathways, cell activities, microenvironment, and mechanical forces [101]. NRP1 is essential for embryonic angiogenesis and vessel remodeling [102]. Understanding these mechanisms is crucial for developing therapeutic strategies that target tumor angiogenesis. Computer simulations can provide insights into blood flow and drug distribution, thereby facilitating effective treatment protocols [101, 103].
Extracellular matrix interactions
Invasion of cancer cells involves interactions between tumor cells and the extracellular matrix (ECM) via various molecular mechanisms. This process involves cytoskeletal dynamics, cell-ECM adhesion, and tissue remodeling in response to microenvironmental signals [104]. Tumor cells use collective and single-cell migration strategies to adapt to microenvironmental challenges [104]. During tumor progression, the ECM undergoes remodeling, with cancer cells and stromal fibroblasts producing matrices and enzymes that support invasion [105]. The tensional force field from cell contraction drives collagen fiber remodeling, which enhances cell polarization and invasion by increasing the matrix stiffness [106]. Cellular contractility is essential because its inhibition reduces tumor cell invasion. The interaction between cellular mechanics and ECM remodeling reveals mechanochemical coupling during invasion [106]. Key molecules involved in invasion include integrins, focal adhesion kinase (FAK), and matrix metalloproteases (MMPs). Integrins mediate cell-ECM adhesion, whereas FAK mediates integrin signalling [107, 108]. MMPs facilitate ECM degradation, and MMP-2 plays a significant role [109]. Cancer-associated fibroblasts contribute to ECM remodeling, promoting tumor rigidity, and FAK activation [110]. Cancer cell invasion involves cellular and molecular mechanisms that enable tumor cells to adapt to microenvironmental conditions and therapeutic challenges.
6. Metastasis Mechanisms
Invasion and migration processes
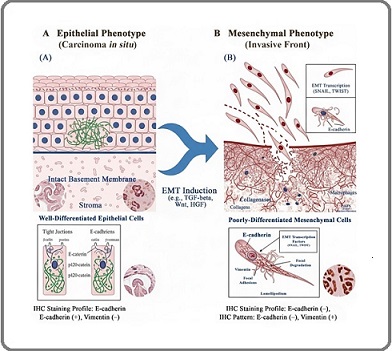
Cancer cell migration and invasion are complex processes involving multiple mechanisms that enable tumor cells to escape from primary sites and metastasize [104,111]. These processes exhibit plasticity and adaptability, allowing cancer cells to respond to microenvironmental cues and overcome therapies [104, 112]. The primary mechanisms include collective, mesenchymal, and amoeboid cell migration, distinguished by cell-cell junctions, actin cytoskeleton organization, matrix adhesion, and protease activity [112]. Cancer cells can transition between these migration modes, demonstrating plasticity that helps them navigate microenvironments and evade therapies [112, 113]. Migration and invasion involve interplay between cellular components and signalling pathways, including cytoskeletal remodeling, ECM degradation, and cell adhesion changes [108, 111]. Cross-talk between the ECM, integrin receptors, matrix metalloproteases, and Rho GTPases modulates cell shape, tension, and migration [108]. The tumor microenvironment influences collective tumor cell invasion [114]. This plasticity manifests itself as histopathological changes. During Epithelial- Mesenchymal Transition (EMT), epithelial cells lose polarity and cell-cell adhesion while gaining mesenchymal markers, thereby driving invasion [115]. This process is visible at tumor ‘invasive fronts.’ Tumor grading reflects this process, with high-grade tumors showing loss of epithelial architecture and a discohesive appearance consistent with EMT, indicating an invasive potential [116] (Figure 1).
Figure 1. Histopathologic and Molecular View of EMT .
Metastatic cascade
Metastatic progression of cancer involves molecular events and cellular transformations that enable tumor cells to spread from primary sites to distant organs. The metastatic cascade begins when cancer cells acquire pro- metastatic characteristics and develop a supportive tumor microenvironment (TME) [117]. This process involves loss of cell polarity, altered adhesion, and deregulated receptor kinase signalling, enabling tumor cell detachment and invasion. Epithelial-to-mesenchymal transition (EMT) is critical, marked by decreased E-cadherin and increased vimentin and Twist1/2 expression (Sridaran et al., 2016). Cancer cells must overcome anoikis and immune surveillance before leaving the primary tumor [117]. The tumor stroma engages in molecular crosstalk within the TME, promoting angiogenesis and immunosuppressive responses. Circulating tumor cells face challenges during their intravasal journey [117]. CTCs interact with the immune system and may be more susceptible to immune effector cells [118]. Successfully extravasating CTCs can colonize distant sites and adapt to survival [117, 119]. This process involves interactions between tumor cells and their microenvironment, creating an immunosuppressive milieu that fosters metastatic tumors [119].
Molecular determinants of metastasis
Metastasis, the leading cause of cancer mortality, involves complex events regulated by molecular factors and signalling pathways. Reactive oxygen species (ROS) signalling facilitates the metastatic cascade and SIRT3 regulates cell migration through ROS control [120]. The invasion-metastasis cascade is influenced by pathways including TGFβ, Wnt/β-Catenin, Hedgehog, p53, and cancer stem cell maintenance, which are modulated by proteins like Kindlin-2 [121]. NF-κB is critical in pancreatic cancer metastasis, regulating cell proliferation, anti-apoptosis, angiogenesis, epithelial-mesenchymal transition, and inflammation [122]. The MAP pathway, regulated by RKIP, contributes to cell growth, invasion, and metastasis [123]. Hypoxia-induced alterations mediated by HIF-1 and pathways, such as c-Src and c-Met, contribute to the metastatic phenotype [124]. It defines that metastasis is regulated by molecular factors and pathways, including ROS signalling, NF-κB, MAP, and hypoxia- induced pathways. Understanding these mechanisms is essential for developing targeted therapies to prevent metastasis. The involvement of tumour stromal cells, particularly endothelial cells, in early tumour metastasis underscores the complexity of this process [125].
Organ-specific metastasis
Cancer cells demonstrate organ-specific preferences during metastasis through interactions between the tumour cells and the target organ microenvironment. Organ tropism is influenced by several mechanisms. The metastatic cascade selectively favours the survival of specific subpopulations of metastatic cells within the primary tumour [126]. These cells have properties that enable organ colonisation. Breast cancer cells exhibit organ-specific malignant behaviour in response to organ- conditioned media, matching their in vivo metastatic potential [127]. The organ microenvironment is crucial for metastatic colonisation. As proposed by Paget’s “seed and soil” hypothesis, the organ microenvironment influences tumour cell invasion, survival, growth, and apoptosis [126]. This interaction occurs via paracrine stimulation by organ-derived growth factors and cytokines [126, 128]. A key insight into this hypothesis is the pre-metastatic niche (PMN), where primary tumours “prepare” distant sites by secreting extracellular vesicles (EVs) that create a favourable microenvironment[129]. Organ-specific metastasis results from the interplay between cancer cell properties and organ microenvironment characteristics. Understanding these mechanisms is crucial for developing targeted therapies and predicting metastasis. Future research should focus on organ-derived factors influencing metastatic and the pre-metastatic niche role [127, 129].
Metastatic colonization
Cancer cells employ mechanisms to colonise distant organs and form secondary tumours. Metastasis involves escape from primary tumours, survival in circulation, and organ colonisation. Cancer stem cells are crucial for initiating new tumours at secondary sites [130]. To colonise organs, cancer cells must overcome shear stress and adapt to new microenvironments [131]. Cancer cells reprogram their metabolism to survive in new environments. This metabolic reprogramming of amino acid metabolism is essential for colonisation of organs such as the lung, liver, brain, peritoneum, and bone [132]. Primary tumours secrete extracellular vesicles (EVs) to form pre-metastatic niches (PMN) that prepare distant sites for colonisation [133]. Metastatic colonisation depends on interactions between cancer cells and the host microenvironment, involving metabolic adaptations and niche formation. Understanding these mechanisms is crucial for developing therapies to prevent metastasis, the main cause of cancer mortality [131-133].
7. Cellular Metabolism in Cancer
Metabolic reprogramming
Cancer cells undergo metabolic reprogramming to facilitate rapid growth and survival in adverse conditions. This metabolic adaptability allows them to fulfil their energy needs and sustain their viability in challenging environments [134]. Key alterations include the Warburg effect and changes in lactate, glutamine, and lipid metabolism to supply intermediates for nucleotide, protein, and fatty acid biosynthesis [135, 136]. Recent research indicates that cancer cells depend on glutamine, lipids, and mitochondrial metabolism [137]. The carnitine system regulates metabolic flexibility, enabling transition between glucose and fatty acid metabolism [134]. MicroRNAs (miRNAs) modulate these metabolic changes and often act as downregulated tumour suppressors in cancer [135, 137]. Cancer cells utilise metabolic reprogramming strategies involving glycolysis, glutaminolysis, lipid metabolism, and mitochondrial function [138, 139]. Understanding these adaptations offers opportunities to develop innovative diagnostic tools and therapeutic strategies [137, 140]. A notable example is the ‘porphyrin overdrive’ mechanism, an imbalanced heme-metabolic pathway representing a promising target for future therapies [141].
Warburg effect
The Warburg effect is a metabolic phenomenon in cancer cells characterised by increased glycolysis and lactate production even in the presence of adequate oxygen [142]. This metabolic reprogramming enables cancer cells to fulfil energy and biosynthetic requirements for rapid proliferation [143].
The Warburg effect includes key metabolic alterations:
1. A preference for aerobic glycolysis over oxidative phosphorylation for ATP production [143].
2. An increase in glucose uptake and lactate production [144].
3. Reprogramming of mitochondrial oxidative phosphorylation [145].
4. Activation of lactate dehydrogenase A to convert pyruvate into lactate [144].
While the Warburg effect is a hallmark of cancer metabolism, its universality remains debated. Some studies show that certain cancer cells exhibit high mitochondrial respiration and low glycolysis [146]. The ‘reverse Warburg effect’ proposes that glycolytic stromal cells secrete lactate, which is used by oxidative cancer cells, highlighting complex metabolic interactions within the TME [146].
The Warburg effect provides advantages to cancer cells:
1. Rapid energy production and increased proliferation [143].
2. Enhanced survival under hypoxic conditions [144].
3. Support for tumor progression and resistance to cell death (Coelho et al., 2018).
4. Promotion of invasiveness and metastasis [144].
These metabolic alterations confer growth advantages to cancer cells and contribute to therapy resistance. Understanding this reprogramming offers opportunities for targeted cancer therapies [142, 144].
Metabolic vulnerabilities
Cancer cells possess metabolic vulnerabilities that can be targeted because of their altered metabolic processes (Table 2).
| Metabolic Pathway | Key Alterations in Cancer | Rationale for Targeting | Example Therapeutic Strategy |
| Glucose Metabolism (Warburg Effect) | Increased aerobic glycolysis | High glucose uptake and l actate production | Glycolysis inhibitors (e.g., 2-DG) MCT1 inhibitors (block lactate export) |
| Glutamine Metabolism | "Glutamine addiction" for biosynthesis | Cancer cells depend on glutamine for nitrogen and anaplerosis | Glutaminase (GLS) inhibitors (e.g., CB-839) |
| Lipid Metabolism | De novo fatty acid synthesis | Required for building new cell membranes | Fatty Acid Synthase ( FASN) inhibitors |
| One-Carbon Metabolism | High demand for folate pathway | Needed for nucleotide (purine/pyrimidine) synthesis | Antifolates (e.g., Methotrexate) |
| Amino Acid Metabolism | Dependency on specific amino acids (e.g., Arginine, Serine) | Some tumors cannot synthesize these "non-essential" amino acids | Arginine-degrading enzymes (Arginase) |
These vulnerabilities arise from metabolic reprogramming during tumor development [147, 148]. A key vulnerability is modified glucose metabolism. Tumors rely on glycolysis for energy production, even in the presence of oxygen (the Warburg effect), which can be exploited with glycolysis inhibitors [149, 150]. The increased lactate production has led to therapies targeting lactate transporters (MCTs) [151]. Another vulnerability is glutamine metabolism dependence, vital for biosynthesis and energy. Glutaminase inhibitors show potential here [149, 150, 152]. Some tumours rely on mitochondrial oxidative phosphorylation (OXPHOS), making them susceptible to OXPHOS inhibitors [149, 151]. Other vulnerabilities include lipid and nucleotide metabolism, as well as specific pathways [147, 153]. Targeting metabolic interactions between cancer cells and tumour microenvironment shows promise [154, 155]. Cancer cells’ metabolic reprogramming offers exploitable vulnerabilities through inhibitors and combination therapies. However, tumour heterogeneity and adaptive resistance require personalised approaches and predictive biomarkers [149, 150].
Metabolic interactions in tumor microenvironment
Metabolic interactions between cancer cells and the tumor microenvironment (TME) are crucial for cancer progression. Cancer cells undergo metabolic reprogramming, altering their glucose and lipid metabolism to adapt to the TME [156, 157]. This reprogramming affects surrounding cells, creating metabolic crosstalk within the TME [66, 158]. Cancer cells stimulate adipocytes to release fatty acids, thereby supporting their growth and survival [159]. These interactions are reciprocal. Cancer-associated fibroblasts (CAFs) and stromal cells influence cancer cell metabolism by supplying nutrients and signals that promote tumor growth and metastasis [160]. Metabolic changes in immune cells within the TME alter their function, potentially enabling cancer cells to evade the immune response [159, 160]. Metabolic interactions between cancer cells and the TME create a supportive environment for tumor growth, metastasis, and therapy resistance. Understanding these interactions offers opportunities to develop therapies targeting metabolic pathways in cancer cells and the TME [156, 157, 160].
8. Molecular Therapeutic Approaches Targeted therapies
Targeted therapies represent a cutting-edge approach to cancer treatment, designed to address molecular and genetic abnormalities that drive tumor growth [161]. These therapies disrupt specific receptor signaling pathways that are crucial for tumor cell proliferation, survival, and migration [162]. Targeted therapies include small-molecule inhibitors, monoclonal antibodies, antibody-drug conjugates, and gene-editing technologies [161]. Their success depends on precise detection and sustained targeting of initial and evolving driver mutations within cancer cells [163]. In non-small cell lung cancer (NSCLC), targeted therapies address specific molecular alterations, such as EGFR mutations, ALK and ROS1 fusions, and BRAF, RET, MET, HER2, and KRAS alterations [164]. This enables personalized treatment, tailoring therapies to a patient’s tumor’s molecular profile [165]. However, challenges persist, including tumor heterogeneity and clonal evolution, leading to drug resistance and treatment failure [163]. While some targeted therapies have shown significant efficacy, others have produced suboptimal outcomes in clinical trials [166]. The development of targeted therapies, driven by advances in molecular profiling techniques, represents progress towards more effective and personalized cancer care [161, 164].
Immunotherapy mechanisms
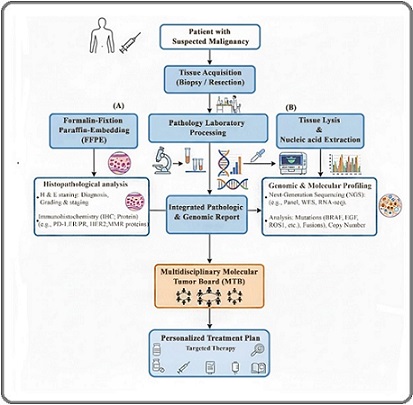
Precision medicine customizes cancer treatments for individual patients by considering their genetic, molecular, and clinical characteristics to deliver more effective therapies while reducing adverse effects. In oncology, precision medicine applies genetic and patient-specific information to optimize preventive and therapeutic care [167]. It uses biomarkers, including genetic, epigenetic, protein, and metabolite markers, to identify patients who are candidates for targeted therapies [168]. This approach has enabled companion diagnostics, where molecular assays measure proteins, genes, or mutations to provide tailored therapy [167]. Patient-derived tumor organoids, combined with genomic sequencing and drug screening, allow researchers to model personalized therapeutic responses [169]. Circulating biomarkers assess patient risk and tumor characteristics and monitor treatment outcomes [170]. Precision medicine aims to complement standard approaches by tailoring treatments to unique tumor characteristics [171]. By integrating genomic profiling and targeted therapies, precision medicine offers the potential to improve cancer treatment outcomes [161, 172]. However, challenges including tumor heterogeneity, limited biomarkers, high costs, and resistance to targeted therapies must be addressed [168] (Figure 2).
Figure 2. Clinical Workflow of Precision Oncology.
Precision medicine
Precision medicine customizes cancer treatments for individual patients by considering their genetic, molecular, and clinical characteristics to deliver more effective therapies while reducing adverse effects. In oncology, precision medicine applies genetic and patient-specific information to optimize preventive and therapeutic care [167]. It uses biomarkers, including genetic, epigenetic, protein, and metabolite markers, to identify patients who are suited for targeted therapies [168]. This approach has enabled companion diagnostics, where molecular assays measure proteins, genes, or mutations to provide tailored therapy [167]. Patient-derived tumor organoids, combined with genomic sequencing and drug screening, enable researchers to model personalized therapeutic responses [169]. Circulating biomarkers assess patient risk and tumor characteristics and monitor treatment outcomes [170]. Precision medicine aims to complement standard approaches by tailoring treatments to unique tumor characteristics [171]. By integrating genomic profiling and targeted therapies, precision medicine offers the potential to improve cancer treatment outcomes [161, 172]. However, challenges including tumor heterogeneity, limited biomarkers, high costs, and resistance to targeted therapies must be addressed [168].
Emerging therapeutic strategies
The latest emerging therapeutic strategies for cancer treatment encompass innovative approaches to overcome drug resistance and enhance patient outcomes. Targeted therapies, combination treatments, and immunotherapies have demonstrated significant potential in addressing cancer’s molecular mechanisms [173]. Nanotechnology-enhanced photodynamic therapy (PDT) with immunotherapy has shown synergistic effects in preclinical studies, enhancing immune responses and mitigating the immunosuppressive tumor microenvironment [174]. Smart therapeutic carriers have emerged as an effective strategy to improve drug delivery and patient outcomes, with promise for cancer immunotherapy. Two-dimensional molybdenum disulfide (MoS2) has demonstrated potential as a platform for cancer theranostics, offering applications in imaging, photothermal therapy, and drug delivery [175]. Emerging therapies for pancreatic cancer include immunotherapy, genetic profiling, and targeted approaches [176]. Targeted therapies and personalized strategies are being investigated to address CRC treatment resistance [177]. Precision medicine and immunotherapy are transforming lung cancer treatment [178]. Cancer therapeutics is rapidly evolving, emphasizing personalized approaches that leverage nanotechnology, immunotherapy, and molecular biology for improved outcomes.
Molecular targeting approaches
Molecular targeting strategies enhance the specificity and efficacy of cancer therapy by focusing on tumor cells’ unique molecular characteristics. These methodologies identify and target specific molecules involved in cancer growth, enabling personalized treatments. Targeted therapies deliver anticancer agents to tumor cells while minimizing damage to adjacent cells, resulting in higher effective doses and improved responses [179]. Aptamers, single-stranded oligonucleotides, bind to target molecules with high specificity, promoting localization and uptake by specific cell populations [179]. Advancements in genomics, proteomics, and transcriptomics have identified molecular targets, leading to the development of monoclonal antibodies and intracellular targeting agents [180]. These therapies inhibit cancer cell proliferation by interfering with tumor growth molecules [181]. Small-molecule inhibitors and antibody-drug conjugates enhance treatment precision [161]. Molecular-targeting approaches enable precise, personalized treatments based on tumors’ genetic and molecular characteristics. This shift from organ-centric to personalized approaches has enabled more effective cancer treatments (Silva et al., 2023; Silva et al., 2024). However, drug resistance, toxicity, and high costs remain challenges to these therapies [181].
In conclusion, cancer is an intricate and evolving disease system shaped by the interplay of genetic mutations, cellular dynamics, and microenvironmental influences. A comprehensive understanding of these molecular mechanisms is essential for developing effective therapeutic strategies. Precision medicine, supported by advances in genomics and molecular profiling, continues to redefine treatment paradigms through targeted and personalized interventions. The integration of artificial intelligence and machine learning into cancer research is accelerating the discovery of predictive biomarkers and therapeutic targets, while patient-derived models such as tumor organoids are enhancing the precision of drug screening and individualized therapy design. Collectively, these emerging approaches signal a transition toward more adaptive, data-driven oncology one that aims not merely to treat but to manage cancer as a chronic, controllable condition.
Declarations
Clinical trial registration
Not applicable.
Conflicts of interest/Competing interests
The authors declare that they have no competing interests.
Availability of data and material
Not applicable. As this is a review article, all data synthesized are from previously published studies, which are fully cited in the reference list.
Code availability
Not applicable.
Authors’ contributions
S.M. contributed to the conception, design, and final drafting of the manuscript. S.M. and J.C. contributed to the literature search and primary drafting of the manuscript.
T.L.S.R. supervised the study. All authors read and approved the final manuscript for submission.
Ethics approval
Not applicable. This article is a review of existing literature and does not involve any new studies on human or animal subjects.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Acknowledgments
The authors wish to acknowledge the management and faculty of St. Xavier’s College (Autonomous), Palayamkottai, for providing the institutional support and academic environment that facilitated this research.
Originality Declaration for Figures
All figures included in this manuscript are original and have been created by the authors specifically for the purposes of this study. No previously published or copyrighted images have been used. The authors confirm that all graphical elements, illustrations, and visual materials were generated from the data obtained in the course of this research or designed uniquely for this manuscript.
Declaration on generative AI and AI-assisted technologies in the writing process
During the preparation of this manuscript, the authors used AI-assisted technologies (Grammarly and Turnitin) for grammar checking, plagiarism detection, and initial outlining.
References
- Epigenetic modulation of cancer stem cells in vitro 3D models Das KJ , Stella JH , Saunders M, et al . Cancer Stem Cells and Signaling Pathways.2024. CrossRef
- Variation of the T Lymphocytes According to Treatment in Breast Cancer Zgura A, Gales L, Bratila E, Mehedintu C, Haineala B, Barac RI , Popa AR , et al . Revista de Chimie.2019;70(5).
- Single-Cell RNA-Sequencing: Opening New Horizons for Breast Cancer Research Xiang L, Rao J, Yuan J, Xie T, Yan H. International Journal of Molecular Sciences.2024;25(17). CrossRef
- Molecular Dimensions of Gastric Cancer: Translational and Clinical Perspectives Choi YY , Noh SH , Cheong J. Journal of Pathology and Translational Medicine.2016;50(1). CrossRef
- Abnormal coagulation and deep venous thrombosis in patients with advanced cancer Johnson M. J., Walker I. D., Sproule M. W., Conkie J.. Clinical and Laboratory Haematology.1999;21(1). CrossRef
- Cell cycle control in cancer Matthews HK , Bertoli C, Bruin RAM . Nature Reviews. Molecular Cell Biology.2022;23(1). CrossRef
- Molecular contribution of BRCA1 and BRCA2 to genome instability in breast cancer patients: review of radiosensitivity assays Sadeghi F, Asgari M, Matloubi M, Ranjbar M, Karkhaneh Yousefi N, Azari T, Zaki-Dizaji M. Biological Procedures Online.2020;22. CrossRef
- Structural basis for inhibition of the cyclin-dependent kinase Cdk6 by the tumour suppressor p16INK4a Russo A. A., Tong L., Lee J. O., Jeffrey P. D., Pavletich N. P.. Nature.1998;395(6699). CrossRef
- The Caulobacter cell cycle: timing, spatial organization and checkpoints Jenal U, Stephens C. Current Opinion in Microbiology.2002;5(6). CrossRef
- Genomic abnormalities and signal transduction dysregulation in malignant mesothelioma cells Sekido Y. Cancer Science.2010;101(1). CrossRef
- Immune vulnerabilities of mutant KRAS in pancreatic cancer Cheng NC , Vonderheide RH . Trends in Cancer.2023;9(11). CrossRef
- The Hallmarks of Cancer Hanahan D, Weinberg RA . Cell.2000;100(1). CrossRef
- Multifaceted Properties of Usnic Acid in Disrupting Cancer Hallmarks Gimła M, Herman-Antosiewicz A. Biomedicines.2024;12(10). CrossRef
- AKT and the Hallmarks of Cancer Sementino E, Hassan D, Bellacosa A, Testa JR . Cancer Research.2024;84(24). CrossRef
- Targeting Six Hallmarks of Cancer in Ovarian Cancer Therapy Gong H, Nie D, Li Z. Current Cancer Drug Targets.2020;20(11). CrossRef
- Transforming growth factor-β and the hallmarks of cancer Tian M, Neil JR , Schiemann WP . Cellular Signalling.2011;23(6). CrossRef
- Ursolic Acid's Alluring Journey: One Triterpenoid vs. Cancer Hallmarks Limami Y, Pinon A, Wahnou H, Oudghiri M, Liagre B, Simon A, Duval RE . Molecules.2023;28(23). CrossRef
- Heat shock proteins as hallmarks of cancer: insights from molecular mechanisms to therapeutic strategies Zuo W, Pang Q, Zhu X, Yang Q, Zhao Q, He G, Han B, Huang W. Journal of Hematology & Oncology.2024;17(1). CrossRef
- Genomic Instability in Cancer: Molecular Mechanisms and Therapeutic Potentials Salmaninejad A, Ilkhani K, Marzban H, Navashenaq JG , Rahimirad S, Radnia F, Yousefi M, et al . Current Pharmaceutical Design.2021;27(28). CrossRef
- Germline mutation contribution to chromosomal instability Chan SH , Ngeow J. Endocrine-Related Cancer.2017;24(9). CrossRef
- High-risk prostate cancer: a disease of genomic instability Tapia-Laliena MA , Korzeniewski N, Hohenfellner M, Duensing S. Urologic Oncology.2014;32(8). CrossRef
- Stop pulling my strings - what telomeres taught us about the DNA damage response Lazzerini-Denchi E, Sfeir A. Nature Reviews. Molecular Cell Biology.2016;17(6). CrossRef
- Effect of ubiquitin protease system on DNA damage response in prostate cancer (Review) Lin Y, Jin X. Experimental and Therapeutic Medicine.2024;27(1). CrossRef
- ESMO recommendations on microsatellite instability testing for immunotherapy in cancer, and its relationship with PD-1/PD-L1 expression and tumour mutational burden: a systematic review-based approach Luchini C., Bibeau F., Ligtenberg M. J. L., Singh N., Nottegar A., Bosse T., Miller R., et al . Annals of Oncology: Official Journal of the European Society for Medical Oncology.2019;30(8). CrossRef
- Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade Le DT , Durham JN , Smith KN , Wang H, Bartlett BR , Aulakh LK , Lu S, et al . Science.2017;357(6349). CrossRef
- Molecular Biology for Cancer Therapy: Review Articles Goshu B. 2021. CrossRef
- Oligonucleotide Arrays for the Detection of ras Mutations Matson RS . Springer Berlin Heidelberg.2000;:724-37. CrossRef
- Tumor suppressor genes: new prospects for cancer research Hollingsworth R. E., Lee W. H.. Journal of the National Cancer Institute.1991;83(2). CrossRef
- Tumor suppressor genes in breast cancer: the gatekeepers and the caretakers Oliveira AM , Ross JS , Fletcher JA . American Journal of Clinical Pathology.2005;124 Suppl. CrossRef
- Id1 overexpression is independent of repression and epigenetic silencing of tumor suppressor genes in melanoma Healey MA , Deaton SL , Alder JK , Winnepenninckx V, Casero RA , Herman JG . Epigenetics.2010;5(5). CrossRef
- Identifying driver mutations from sequencing data of heterogeneous tumors in the era of personalized genome sequencing Zhang J, Liu J, Sun J, Chen C, Foltz G, Lin B. Briefings in Bioinformatics.2014;15(2). CrossRef
- Passenger mutations can accelerate tumor suppressor gene inactivation in cancer evolution [Preprint] Wodarz D, Newell AC , Komarova NL . Cold Spring Harbor Laboratory.2017. CrossRef
- Passenger mutations can accelerate tumour suppressor gene inactivation in cancer evolution Wodarz D, Newell AC , Komarova NL . Journal of the Royal Society, Interface.2018;15(143). CrossRef
- Tug-of-war between driver and passenger mutations in cancer and other adaptive processes McFarland CD , Mirny LA , Korolev KS . Proceedings of the National Academy of Sciences of the United States of America.2014;111(42). CrossRef
- The Damaging Effect of Passenger Mutations on Cancer Progression McFarland CD , Yaglom JA , Wojtkowiak JW , Scott JG , Morse DL , Sherman MY , Mirny LA . Cancer Research.2017;77(18). CrossRef
- Identification of constrained cancer driver genes based on mutation timing Sakoparnig T, Fried P, Beerenwinkel N. PLoS computational biology.2015;11(1). CrossRef
- DNA methylation and gene silencing in cancer Baylin SB . Nature Clinical Practice. Oncology.2005;2 Suppl 1. CrossRef
- Epigenetics in cancer: targeting chromatin modifications Ellis L, Atadja PW , Johnstone RW . Molecular Cancer Therapeutics.2009;8(6). CrossRef
- Histone modification and personalized epigenomics in cancer Ramazi S, Dadzadi M, Sahafnejad Z, et al . Personalized Epigenetics.2024. CrossRef
- The language of chromatin modification in human cancers Zhao S, Allis CD , Wang GC . Nature Reviews. Cancer.2021;21(7). CrossRef
- Characterization of Chromatin Structure-associated Histone Modifications in Breast Cancer Cells Hong CO , Choe MK , Roh T. Genomics & Informatics.2012;10(3). CrossRef
- The crosstalk between m6A RNA methylation and other epigenetic regulators: a novel perspective in epigenetic remodeling Zhao Y, Chen Y, Jin M, Wang J. Theranostics.2021;11(9). CrossRef
- Genomic landscape of diffuse glioma revealed by whole genome sequencing Kinnersley B, Jung J, Cornish AJ Alex J., Chubb D, Laxton R, Frangou A, Gruber AJ , et al . Nature Communications.2025;16(1). CrossRef
- An integrated analysis of the epigenetic, genetic, and transcriptional patterns associated with outcome across cancer types Smith JC , Sheltzer JM . 2017. CrossRef
- Whole-genome sequencing and comprehensive molecular profiling identify new driver mutations in gastric cancer Wang K, Yuen ST , Xu J, Lee SP , Yan HHN , Shi ST , Siu HC , et al . Nature Genetics.2014;46(6). CrossRef
- A multilevel pan-cancer map links gene mutations to cancer hallmarks Knijnenburg TA , Bismeijer T, Wessels LFA , Shmulevich I. Chinese Journal of Cancer.2015;34(10). CrossRef
- The molecular tumor board—a key element of precision oncology Boos L, Wicki A. memo - Magazine of European Medical Oncology.2024;17(3). CrossRef
- Protein kinase signaling networks in cancer Brognard J, Hunter T. Current Opinion in Genetics & Development.2011;21(1). CrossRef
- The ERK cascade as a prototype of MAPK signaling pathways Rubinfeld H, Seger R. Methods in Molecular Biology.2004;250. CrossRef
- Ras-MAP kinase signaling pathways and control of cell proliferation: relevance to cancer therapy Shapiro P. Critical Reviews in Clinical Laboratory Sciences.2002;39(4-5). CrossRef
- Cyclic AMP selectively uncouples mitogen-activated protein kinase cascades from activating signals Pearson GW , Earnest S, Cobb MH . Molecular and Cellular Biology.2006;26(8). CrossRef
- Molecular Targeting of Cell Death Signal Transduction Pathways in Cancer Petak I, Houghton JA , Kopper L. Current Signal Transduction Therapy.2006;1(1). CrossRef
- Current strategies to target the anti-apoptotic Bcl-2 protein in cancer cells Osford SME , Dallman CL , Johnson PWM , Ganesan A., Packham G. Current Medicinal Chemistry.2004;11(8). CrossRef
- DNA methylation and apoptosis resistance in cancer cells Hervouet E, Cheray M, Vallette FM , Cartron P. Cells.2013;2(3). CrossRef
- Targeting endocytosis to sensitize cancer cells to programmed cell death Chan ET , Kural C. Biochemical Society Transactions.2024;52(4). CrossRef
- Cell death pathologies: targeting death pathways and the immune system for cancer therapy Pentimalli F, Grelli S, Di Daniele N, Melino G, Amelio I. Genes and immunity.2018. CrossRef
- Anti-apoptotic mechanisms of drug resistance in cancer Wilson T. R., Johnston P. G., Longley D. B.. Current Cancer Drug Targets.2009;9(3). CrossRef
- Nanoluciferase-based complementation assays to monitor activation, modulation and signaling of receptor tyrosine kinases (RTKs) Dosquet H, Neirinckx V, Meyrath M, Wantz M, Haan S, Niclou SP , Szpakowska M, Chevigné A. Methods in Enzymology.2023;682. CrossRef
- Receptor tyrosine kinases: mechanisms of activation and signaling Hubbard SR , Miller WT . Current Opinion in Cell Biology.2007;19(2). CrossRef
- Epidermal growth factor receptor family activation and intracellular signalling. Northeastern University [Dissertation] Richman JM . . CrossRef
- Receptor tyrosine kinase signaling: a view from quantitative proteomics Dengjel J, Kratchmarova I, Blagoev B. Molecular bioSystems.2009;5(10). CrossRef
- Receptor tyrosine kinases (RTKs) in breast cancer: signaling, therapeutic implications and challenges Butti R, Das S, Gunasekaran VP , Yadav AS , Kumar D, Kundu GC . Molecular Cancer.2018;17(1). CrossRef
- Growth Factor Receptor Tyrosine Kinase Inhibitors in Non-small cell lung cancer Manzo LM , Lawaly M, Yu L. Indian Journal of Pharmaceutical and Biological Research.2015;3(02). CrossRef
- Colorectal cancer: Metabolic interactions reshape the tumor microenvironment Chen J, Zhu H, Yin Y, Jia S, Luo X. Biochimica Et Biophysica Acta. Reviews on Cancer.2022;1877(5). CrossRef
- EV-miRNA-Mediated Intercellular Communication in the Breast Tumor Microenvironment Sepúlveda F, Mayorga-Lobos C, Guzmán K, Durán-Jara E, Lobos-González . International Journal of Molecular Sciences.2023;24(17). CrossRef
- Crosstalk between extracellular vesicles and tumor-associated macrophage in the tumor microenvironment Chen QY , Gao B, Tong D, Huang C. Cancer Letters.2023;552. CrossRef
- Tunneling Nanotubes: Intercellular Conduits for Direct Cell-to-Cell Communication in Cancer Lou E, Subramanian Su. Intercellular Communication in Cancer.2015. CrossRef
- Decoding the Complexity of Immune-Cancer Cell Interactions: Empowering the Future of Cancer Immunotherapy Maffuid K, Cao Y. Cancers.2023;15(16). CrossRef
- Dissecting intercellular signaling with mass spectrometry-based proteomics Swietlik JJ , Sinha A, Meissner F. Current Opinion in Cell Biology.2020;63. CrossRef
- Investigation of cellular communication and signaling pathways in tumor microenvironment for high TP53-expressing osteosarcoma cells through single-cell RNA sequencing Xiong K, Fang Y, Qiu B, Chen C, Huang N, Liang F, Huang C, et al . Medical Oncology.2024;41(5). CrossRef
- Intracellular and intercellular signaling networks in cancer initiation, development and precision anti-cancer therapy: RAS acts as contextual signaling hub Csermely P, Korcsmáros T, Nussinov R. Seminars in Cell & Developmental Biology.2016;58. CrossRef
- Overcoming Resistance Mechanisms to Immune Checkpoint Inhibitors: Leveraging the Anti-Tumor Immune Response Coschi CH , Juergens RA . Current Oncology.2023;31(1). CrossRef
- Bypassing anti-PD-(L)1 therapy: Mechanisms and management strategies Mortezaee K, Majidpoor J, Najafi S, Tasa D. Biomedicine & Pharmacotherapy = Biomedecine & Pharmacotherapie.2023;158. CrossRef
- Overcoming acquired resistance to cancer immune checkpoint therapy: potential strategies based on molecular mechanisms Wang B, Han Y, Zhang Y, Zhao Q, Wang H, Wei J, Meng L, Xin Y, Jiang X. Cell & Bioscience.2023;13(1). CrossRef
- TREM2: Keeping Pace With Immune Checkpoint Inhibitors in Cancer Immunotherapy Qiu H, Shao Z, Wen X, Jiang J, Ma Q, Wang Y, Huang L, Ding X, Zhang L. Frontiers in Immunology.2021;12. CrossRef
- Communication between EMT and PD-L1 signaling: New insights into tumor immune evasion Jiang Y, Zhan H. Cancer Letters.2020;468. CrossRef
- Cancer Cell Metabolism Bolsters Immunotherapy Resistance by Promoting an Immunosuppressive Tumor Microenvironment Jiang Z, Hsu JL , Li Y, Hortobagyi GN , Hung M. Frontiers in Oncology.2020;10. CrossRef
- Overcoming cold tumors: a combination strategy of immune checkpoint inhibitors Ouyang P, Wang L, Wu J, Tian Y, Chen C, Li D, Yao Z, et al . Frontiers in Immunology.2024;15. CrossRef
- Immune-microenvironment dynamics in cancer: Implications for immunotherapy Turupati P, Usa D, India D, et al . Int J Adv Res.2024;12(05):903-907. CrossRef
- Single-cell RNA sequencing reveals the landscape of the cellular ecosystem of primary hepatocellular carcinoma Yin Z, Song Y, Wang L. Cancer Cell International.2024;24(1). CrossRef
- Review on Tumour Microenvironment Cell Types Associated with Metastatic Cancer Choudhary SG , Potdar PD . Diseases & Research.2023;3(2). CrossRef
- Unraveling the intricacies of cancer-associated fibroblasts: a comprehensive review on metabolic reprogramming and tumor microenvironment crosstalk Ahuja S, Sureka N, Zaheer S. APMIS: acta pathologica, microbiologica, et immunologica Scandinavica.2024;132(12). CrossRef
- Extracellular matrix-mediated cellular communication in the heart Valiente-Alandi I, Schafer AE , Blaxall BC . Journal of Molecular and Cellular Cardiology.2016;91. CrossRef
- Cancer cell plasticity: from cellular, molecular, and genetic mechanisms to tumor heterogeneity and drug resistance Bhat GR , Sethi I, Sadida HQ , Rah B, Mir R, Algehainy N, Albalawi IA , et al . Cancer Metastasis Reviews.2024;43(1). CrossRef
- Matrix Metalloproteinases Shape the Tumor Microenvironment in Cancer Progression Niland S, Riscanevo AX , Eble JA . International Journal of Molecular Sciences.2021;23(1). CrossRef
- Extracellular Matrix Influencing HGF/c-MET Signaling Pathway: Impact on Cancer Progression Noriega-Guerra H, Freitas VM . International Journal of Molecular Sciences.2018;19(11). CrossRef
- Cell-to-cell contact-mediated regulation of tumor behavior in the tumor microenvironment Sato A, Rahman NIA , Shimizu A, Ogita H. Cancer Science.2021;112(10). CrossRef
- Insidious changes in stromal matrix fuel cancer progression Miles FL , Sikes RA . Molecular cancer research: MCR.2014;12(3). CrossRef
- Gap junction as an intercellular glue: Emerging roles in cancer EMT and metastasis Mao X, Li Q, Gao Y, Zhou H, Liu Z, Jin W. Cancer Letters.2016;381(1). CrossRef
- Mesenchymal stromal cells as cancer promoters Antoon R, Overdevest N, Saleh AH , Keating A. Oncogene.2024;43(49). CrossRef
- Tumor microenvironment and cancer metastasis: molecular mechanisms and therapeutic implications Biray Avci C, Goker Bagca B, Nikanfar M, Takanlou LS , Takanlou MS , Nourazarian A. Frontiers in Pharmacology.2024;15. CrossRef
- Chronic inflammation and cytokines in the tumor microenvironment Landskron G, De la Fuente M, Thuwajit P, Thuwajit C, Hermoso MA . Journal of Immunology Research.2014;2014. CrossRef
- Crosstalk Between Inflammatory Signaling and Methylation in Cancer Das D, Karthik N, Taneja R. Frontiers in Cell and Developmental Biology.2021;9. CrossRef
- Promotion of breast cancer development via Th2 polarization by TSLP (165.2) Xu K, Pedroza-Gonzalez A, Wu T, Aspord C, Shafer S, Marches F, Gallegos M, et al . The Journal of Immunology.2011;186(1_Supplement). CrossRef
- IL-6 in inflammation, autoimmunity and cancer Hirano T. International Immunology.2021;33(3). CrossRef
- Old and New Players of Inflammation and Their Relationship With Cancer Development Chavez-Dominguez R, Perez-Medina M, Aguilar-Cazares D, Galicia-Velasco M, Meneses-Flores M, Islas-Vazquez L, Camarena A, Lopez-Gonzalez JS . Frontiers in Oncology.2021;11. CrossRef
- Insight on the cellular and molecular basis of blood vessel formation: A specific focus on tumor targets and therapy Stephen NM , Deepika UR , Maradagi T, Sugawara T, Hirata T, Ganesan P. MedComm – Oncology.2023;2(1). CrossRef
- Extracellular vesicles: emerging mediators of intercellular communication and tumor angiogenesis Yamada NO . Annals of Translational Medicine.2017;5(3). CrossRef
- Role of hypoxia in tumor angiogenesis-molecular and cellular angiogenic crosstalk Acker T, Plate KH . Cell and Tissue Research.2003;314(1). CrossRef
- The Angiogenic Switch in Tumorigenesis Wicki A, Christofori G. Tumor Angiogenesis: Basic Mechanisms and Cancer Therapy.2008. CrossRef
- Integrative models of vascular remodeling during tumor growth Rieger H, Welter M. Wiley Interdisciplinary Reviews. Systems Biology and Medicine.2015;7(3). CrossRef
- Neuropilin regulation of angiogenesis, arteriogenesis, and vascular permeability Plein A, Fantin A, Ruhrberg C. Microcirculation.2014;21(4). CrossRef
- A computational model predicting disruption of blood vessel development Kleinstreuer N, Dix D, Rountree M, Baker N, Sipes N, Reif D, Spencer R, Knudsen T. PLoS computational biology.2013;9(4). CrossRef
- Plasticity of Cancer Cell Invasion-Mechanisms and Implications for Therapy Te Boekhorst V., Friedl P.. Advances in Cancer Research.2016;132. CrossRef
- The Role of Extracellular Matrix Remodeling in Skin Tumor Progression and Therapeutic Resistance Fromme JE , Zigrino P. Frontiers in Molecular Biosciences.2022;9. CrossRef
- Tensile force field plays a crucial role in local invasion of tumor cells through a mechano-chemical coupling mechanism Meng J, Xu X, Jiang C, Xia P, Xu P, Tian L, Xu Y, et al . Soft Matter.2024;20(30). CrossRef
- Noncanonical Wnt/Ror2 signaling regulates cell-matrix crosstalk to prompt directional tumor cell invasion and dissemination in breast cancer Si H, Zhao N, Pedroza A, Rosen JM , Creighton CJ , Roarty K. 2021. CrossRef
- Mechanisms of Tumor Cell Migration and Invasion in Lung Cancer Metastasis Thodeti CK , Ghosh K. Lung Cancer Metastasis: Novel Biological Mechanisms and Impact on Clinical Practice.2010. CrossRef
- Extracellular matrix 6: role of matrix metalloproteinases in tumor invasion and metastasis Stetler-Stevenson W. G., Liotta L. A., Kleiner D. E.. FASEB journal: official publication of the Federation of American Societies for Experimental Biology.1993;7(15). CrossRef
- Hic-5 remodeling of the stromal matrix promotes breast tumor progression Goreczny G. J., Ouderkirk-Pecone J. L., Olson E. C., Krendel M., Turner C. E.. Oncogene.2017;36(19). CrossRef
- Redox regulation of cancer cell migration and invasion Tochhawng L, Deng S, Pervaiz S, Yap CT . Mitochondrion.2013;13(3). CrossRef
- Plasticity of cancer cell invasion: Patterns and mechanisms Wu J, Jiang J, Chen B, Wang K, Tang Y, Liang X. Translational Oncology.2021;14(1). CrossRef
- Tumour-cell invasion and migration: diversity and escape mechanisms Friedl P, Wolf K. Nature Reviews. Cancer.2003;3(5). CrossRef
- The role of tumor microenvironment in collective tumor cell invasion Wu J, Sheng S, Liang X, Tang Y. Future Oncology.2017;13(11). CrossRef
- EMT in cancer Brabletz T, Kalluri R, Nieto MA , Weinberg RA . Nature Reviews. Cancer.2018;18(2). CrossRef
- The basics of epithelial-mesenchymal transition Kalluri R, Weinberg RA . The Journal of Clinical Investigation.2009;119(6). CrossRef
- Cancer metastasis: Molecular mechanisms and clinical perspectives Khan SU , Fatima K, Malik F, Kalkavan H, Wani A. Pharmacology & Therapeutics.2023;250. CrossRef
- Crucial roles of circulating tumor cells in the metastatic cascade and tumor immune escape: biology and clinical translation Pantel K, Alix-Panabières C. Journal for Immunotherapy of Cancer.2022;10(12). CrossRef
- The complexity of immune evasion mechanisms throughout the metastatic cascade Haynes NM , Chadwick TB , Parker BS . Nature Immunology.2024;25(10). CrossRef
- Inhibition of epithelial cell migration and Src/FAK signaling by SIRT3 Lee JJ , Ven RAH , Zaganjor E, Ng MR , Barakat A, Demmers JJPG , Finley LWS , et al . Proceedings of the National Academy of Sciences of the United States of America.2018;115(27). CrossRef
- Role of Kindlin-2 in cancer progression and metastasis Wang W, Kansakar Ur, Markovic V, Sossey-Alaoui K. Annals of Translational Medicine.2020;8(14). CrossRef
- Roles of NF-κB signaling pathway in regulatory network of pancreatic cancer metastasis DU Y, YOU L, REN X, LIU Z, ZHAO Y. International Journal of Surgery.2014.
- Identification of novel metastasis suppressor signaling pathways for breast cancer Minn AJ , Bevilacqua E, Yun J, Rosner MR . Cell Cycle.2012;11(13). CrossRef
- Impact of Tumor Hypoxia, Src, and Met Signaling in the Dissemination of Tumor Cells Dai Y, Shi W, Molnar N, Siemann D. 2011. CrossRef
- Tumor endothelial cells accelerate tumor metastasis Maishi N, Hida K. Cancer Science.2017;108(10). CrossRef
- Molecular Mechanisms of Tumor Cell Survival, Growth, and Apoptosis During Organ-Specific Metastasis Radinsky R., Bucana C. D., Cho C. H.. Microscopy and Microanalysis.1997;3(S2).
- Generation of Organ-conditioned Media and Applications for Studying Organ-specific Influences on Breast Cancer Metastatic Behavior Piaseczny MM , Pio GM , Chu JE , Xia Y, Nguyen K, Goodale D, Allan A. Journal of Visualized Experiments: JoVE.2016;(112). CrossRef
- Paracrine growth regulation of human colon carcinoma organ-specific metastasis Radinsky R.. Cancer Metastasis Reviews.1993;12(3-4). CrossRef
- Multi-stage mechanisms of tumor metastasis and therapeutic strategies Liu Z, Chen J, Ren Y, Liu S, Ba Y, Zuo A, Luo P, et al . Signal Transduction and Targeted Therapy.2024;9(1). CrossRef
- An Intrasplenic Injection Model for the Study of Cancer Stem Cell Seeding Capacity Dafflon C, Santamaría-Martínez A, Ordóñez-Morán P. Methods in Molecular Biology.2020;2171. CrossRef
- Metastatic colonization: settlement, adaptation and propagation of tumor cells in a foreign tissue environment Shibue T, Weinberg RA . Seminars in Cancer Biology.2011;21(2). CrossRef
- Amino acid metabolic reprogramming in tumor metastatic colonization Wang Z, Wu X, Chen H, Wang K. Frontiers in Oncology.2023;13. CrossRef
- Formation of pre-metastatic niches induced by tumor extracellular vesicles in lung metastasis Liu Y, Mao D, Wang H, Che X, Chen Y. Pharmacological Research.2023;188. CrossRef
- The carnitine system and cancer metabolic plasticity Melone MAB , Valentino A, Margarucci S, Galderisi U, Giordano A, Peluso G. Cell Death & Disease.2018;9(2). CrossRef
- MicroRNAs as regulators of tumor metabolism Ruggieri F, Jonas K, Ferracin M, Dengler M, Jӓger V, Pichler M. Endocrine-Related Cancer.2023;30(8). CrossRef
- The Metabolic Landscape of Lung Cancer: New Insights in a Disturbed Glucose Metabolism Vanhove K, Graulus G, Mesotten L, Thomeer M, Derveaux E, Noben J, Guedens W, Adriaensens P. Frontiers in Oncology.2019;9. CrossRef
- MicroRNAs as Regulators of Cancer Cell Energy Metabolism Suriya Muthukumaran N, Velusamy P, Akino Mercy CS , Langford D, Natarajaseenivasan K, Shanmughapriya S. Journal of Personalized Medicine.2022;12(8). CrossRef
- Tumor metabolic reprogramming in lung cancer progression Li X, Liu M, Liu H, Chen J. Oncology Letters.2022;24(2). CrossRef
- Shutting off the fuel supply to target metabolic vulnerabilities in multiple myeloma Rana PS , Goparaju K, Driscoll JJ . Frontiers in Oncology.2023;13. CrossRef
- Biomarker selection and imaging design in cancer: A link with biochemical pathways for imminent engineering Ali JS , Ain NU , Naz S, Zia M. Heliyon.2020;6(2). CrossRef
- Porphyrin overdrive rewires pan-cancer cell metabolism Adapa SR , Hunter GA , Amin NE , Marinescu C, Borsky A, Sagatys EM , Sebti SM , et al . 2023. CrossRef
- Lactoferrin conjugated radicicol nanoparticles enhanced drug delivery and cytotoxicity in prostate cancer cells Kooshan Z, Srinivasan S, Janjua TI , Popat A, Batra J. European Journal of Pharmacology.2025;991. CrossRef
- Pathogens Hijack Host Cell Metabolism: Intracellular Infection as a Driver of the Warburg Effect in Cancer and Other Chronic Inflammatory Conditions Proal AD , VanElzakker MB . Immunometabolism.2021;3(1). CrossRef
- Contemporary Perspectives on the Warburg Effect Inhibition in Cancer Therapy Kozal K, Jóźwiak P, Krześlak A. Cancer Control: Journal of the Moffitt Cancer Center.2021;28. CrossRef
- Metabolic reprogramming in glioblastoma: the influence of cancer metabolism on epigenetics and unanswered questions Agnihotri S, Zadeh G. Neuro-Oncology.2016;18(2). CrossRef
- Metabolic interplay between glycolysis and mitochondrial oxidation: The reverse Warburg effect and its therapeutic implication Lee M. World Journal of Biological Chemistry.2015;6(3). CrossRef
- Therapeutic strategies targeting metabolic characteristics of cancer cells Bai R, Meng Y, Cui J. Critical Reviews in Oncology/Hematology.2023;187. CrossRef
- Oncogenic regulation of tumor metabolic reprogramming Tarrado-Castellarnau M, Atauri P, Cascante M. Oncotarget.2016;7(38). CrossRef
- Targeting cancer metabolism through synthetic lethality-based combinatorial treatment strategies Bajpai R, Shanmugam M. Current Opinion in Oncology.2018;30(5). CrossRef
- Metabolic vulnerabilities in cancer: A new therapeutic strategy El-Tanani M, Rabbani SA , El-Tanani Y, Matalka II . Critical Reviews in Oncology/Hematology.2024;201. CrossRef
- Therapeutic Targeting of Tumor Cells and Tumor Immune Microenvironment Vulnerabilities Kalyanaraman B, Cheng G, Hardy M. Frontiers in Oncology.2022;12. CrossRef
- The role of glutamine metabolism in castration-resistant prostate cancer Zhao B, Wang J, Chen L, Wang H, Liang C, Huang J, Xu L. Asian Journal of Andrology.2023;25(2). CrossRef
- Targeting metabolic vulnerabilities of cancer: Small molecule inhibitors in clinic Tripathi SC , Fahrmann JF , Vykoukal JV , Dennison JB , Hanash SM . Cancer Reports.2019;2(1). CrossRef
- Effects of metabolic cancer therapy on tumor microenvironment Hyroššová P, Milošević M, Škoda J, Vachtenheim J, Rohlena J, Rohlenová K. Frontiers in Oncology.2022;12. CrossRef
- Targeting Metabolism in Cancer Cells and the Tumour Microenvironment for Cancer Therapy Li J, Eu JQ , Kong LR , Wang L, Lim YC , Goh BC , Wong ALA . Molecules.2020;25(20). CrossRef
- Metabolic Switch in the Tumor Microenvironment Determines Immune Responses to Anti-cancer Therapy Wegiel B, Vuerich M, Daneshmandi S, Seth P. Frontiers in Oncology.2018;8. CrossRef
- The Tumor Microenvironment: Signal Transduction Zhang X, Ma H, Gao Y, Liang Y, Du Y, Hao S, Ni T. Biomolecules.2024;14(4). CrossRef
- Metabolic crosstalk in the breast cancer microenvironment Dias AS , Almeida CR , Helguero LA , Duarte IF . European Journal of Cancer.2019;121. CrossRef
- Lipids in the tumor microenvironment: From cancer progression to treatment Corn KC , Windham MA , Rafat M. Progress in Lipid Research.2020;80. CrossRef
- Metabolic reprogramming and crosstalk of cancer-related fibroblasts and immune cells in the tumor microenvironment Zhu Y, Li X, Wang L, Hong X, Yang J. Frontiers in Endocrinology.2022;13. CrossRef
- Advances in Targeted Therapy for Cancer: Precision Medicine Approaches Asha BB . IDOSR J Appl Sci.2024;9(2):63-7. CrossRef
- Targeted therapies for solid tumors: current status and future perspectives Stoffel A. BioDrugs: Clinical Immunotherapeutics, Biopharmaceuticals and Gene Therapy.2010;24(5). CrossRef
- Genetic Heterogeneity and Clonal Evolution of Tumor Cells and their Impact on Precision Cancer Medicine Sabaawy HE . Journal of Leukemia.2013;1(4). CrossRef
- Targeted Therapy for Non-Small Cell Lung Cancer Noor ZS, , Cummings AL , Spiegel ML , et al . Semin Respir Crit Care Med.2020;41(3):409-434. CrossRef
- Identification of driver mutations in lung cancer: first step in personalized cancer Planchard D. Targeted Oncology.2013;8(1). CrossRef
- Genomic Alterations and Targeted Therapy in Gastric and Esophageal Adenocarcinoma Pectasides E. Clinical Therapeutics.2016;38(7). CrossRef
- Precision and personalized medicine: new way for cancer management Bhambre AS , Giri AB , Mundhe SA , Mahadik AB , Bhambre AS , Giri AB , Mundhe SA , Mahadik AB . GSC Biological and Pharmaceutical Sciences.2022;18(3). CrossRef
- Unlocking the power of precision medicine: exploring the role of biomarkers in cancer management Gadade DD , Jha H, Kumar C, Khan F. Future Journal of Pharmaceutical Sciences.2024;10(1). CrossRef
- Abstract A20: Personalized models to guide precision medicine Pauli C, Puca L, Hopkins B, Emerling BM , Sboner A, Elemento O, McNary TJ , et al . Clinical Cancer Research.2016;22(16_Supplement). CrossRef
- Advancing Precision Medicine: Tailoring Personalized Therapies Matchett KB , Lynam-Lennon N, Brown JAL , Watson RW . Jenny Stanford.2019;:393-410. CrossRef
- A Review on Patient-derived 3D Micro Cancer Approach for Drug Screen in Personalized Cancer Medicine Sekeroglu ZA , Sekeroglu V. Current Cancer Drug Targets.2025;25(2). CrossRef
- Emerging Precision Medicine Approaches for Lung Neuroendocrine Tumors Mulvey CK . Cancers.2023;15(23). CrossRef
- Emerging Therapeutic Strategies to Overcome Drug Resistance in Cancer Cells Garg P, Malhotra J, Kulkarni P, Horne D, Salgia R, Singhal SS . Cancers.2024;16(13). CrossRef
- Revolutionizing cancer treatment: nanotechnology-enabled photodynamic therapy and immunotherapy with advanced photosensitizers Jia J, Wu X, Long G, Yu J, He W, Zhang H, Wang D, Ye Z, Tian J. Frontiers in Immunology.2023;14. CrossRef
- Recent developments in two-dimensional molybdenum disulfide-based multimodal cancer theranostics Yu X, Xu C, Sun J, Xu H, Huang H, Gan Z, George A, Ouyang S, Liu F. Journal of Nanobiotechnology.2024;22(1). CrossRef
- Molecular underpinnings of pancreatic cancer: Innovations in treatment and future perspectives Tez M, Sahingöz E, Martlı F. Gastrointestinal Tract.2023;1. CrossRef
- Chapter Six - Emerging treatment approaches for colorectal cancer treatment resistance. In: Colorectal Cancer Treatment Resistance. Mabonga L, Fourie L, Kappo AP , et al . Elsevier.2025. CrossRef
- Cutting-Edge Therapies for Lung Cancer La'ah AS , Chiou S. Cells.2024;13(5). CrossRef
- Aptamer-Functionalized Nanoparticles as "Smart Bombs": The Unrealized Potential for Personalized Medicine and Targeted Cancer Treatment Benedetto G, Vestal CG , Richardson C. Targeted Oncology.2015;10(4). CrossRef
- Molecular basis and clinical application of targeted therapy in oncology Tilak T. V. S. V. G. K., Patel A, Kapoor A. Medical Journal, Armed Forces India.2023;79(2). CrossRef
- Limitations of molecularly targeted therapy Balik K, Bajek A, Modrakowska P, Maj M, Kaźmierski Ł.. Med Res J.2019;4(2):99-105. CrossRef
Content
Abstract 1. Introduction 2. Cellular Foundations of Cancer 3. Genetic and Molecular Basis of Cancer 4. Molecular Signalling in Cancer Development 5. Cancer Microenvironment 6. Metastasis Mechanisms 7. Cellular Metabolism in Cancer 8. Molecular Therapeutic Approaches Targeted therapies Acknowledgments ReferencesLicense

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Copyright
© Asian Pacific Journal of Cancer Care , 2026
Author Details
How to Cite
- Abstract viewed - 0 times
- PDF (FULL TEXT) downloaded - 0 times
- XML downloaded - 0 times